Probe CUST_44879_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44879_PI426222305 | JHI_St_60k_v1 | DMT400044098 | GTATGGGATACCAACATCAAGTATTCTCAAGGCAGGGGCATTGGCATTGTTGTTAGTCCA |
All Microarray Probes Designed to Gene DMG400017110
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44879_PI426222305 | JHI_St_60k_v1 | DMT400044098 | GTATGGGATACCAACATCAAGTATTCTCAAGGCAGGGGCATTGGCATTGTTGTTAGTCCA |
| CUST_44882_PI426222305 | JHI_St_60k_v1 | DMT400044097 | GTATGGGATACCAACATCAAGTATTCTCAAGGCAGGGGCATTGGCATTGTTGTTAGTCCA |
Microarray Signals from CUST_44879_PI426222305
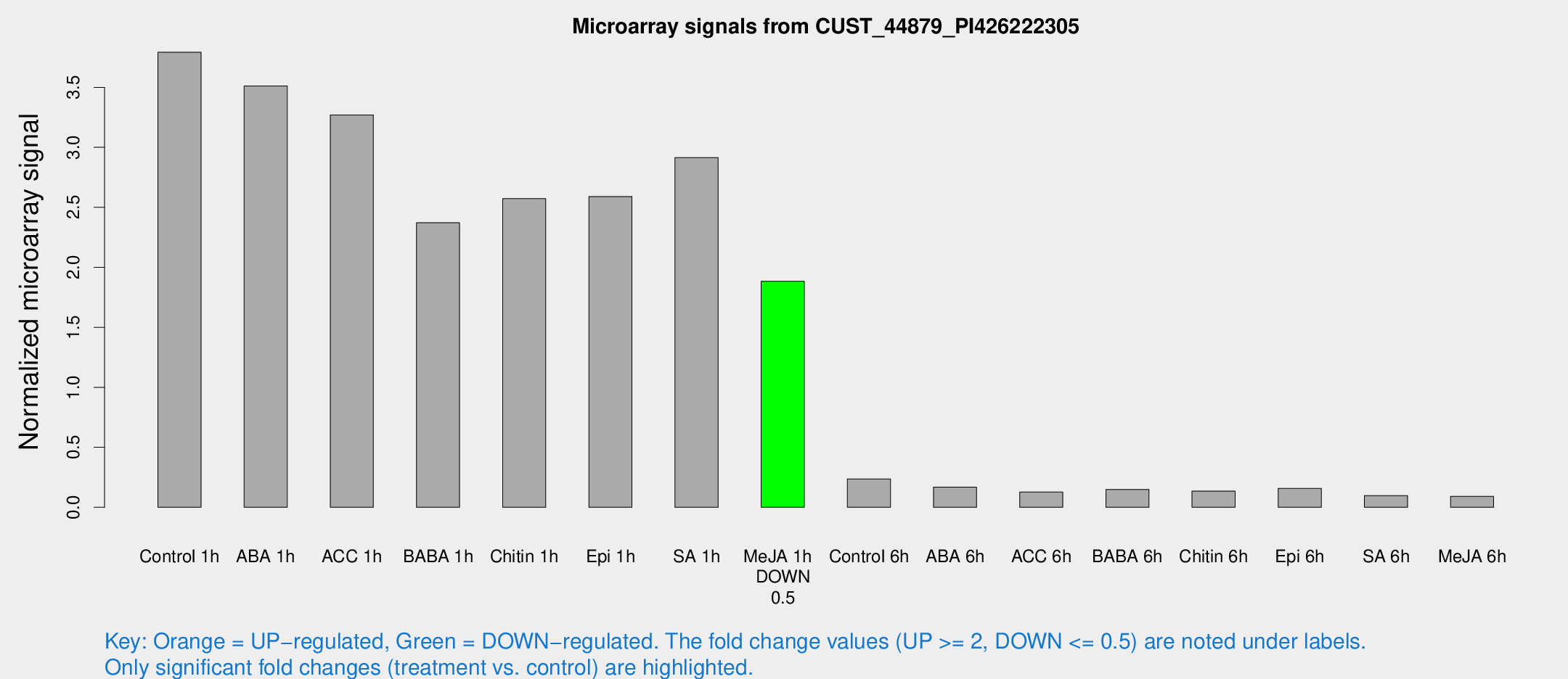
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 10838.9 | 1432.12 | 3.79249 | 0.25703 |
| ABA 1h | 8752.91 | 523.227 | 3.51124 | 0.289295 |
| ACC 1h | 9534.45 | 979.048 | 3.26994 | 0.188794 |
| BABA 1h | 6874.77 | 1603.35 | 2.37172 | 0.414365 |
| Chitin 1h | 6615.2 | 930.176 | 2.57209 | 0.379119 |
| Epi 1h | 6298.35 | 363.907 | 2.58998 | 0.149538 |
| SA 1h | 8456.64 | 664.526 | 2.91462 | 0.168279 |
| Me-JA 1h | 4351.34 | 375.721 | 1.88354 | 0.108755 |
| Control 6h | 714.475 | 171.292 | 0.236931 | 0.0433618 |
| ABA 6h | 511.483 | 65.1096 | 0.168645 | 0.0170528 |
| ACC 6h | 428.666 | 95.6482 | 0.127136 | 0.00911903 |
| BABA 6h | 476.222 | 56.5524 | 0.149292 | 0.022339 |
| Chitin 6h | 415.635 | 69.8174 | 0.135423 | 0.0237839 |
| Epi 6h | 530.915 | 140.195 | 0.156944 | 0.0466095 |
| SA 6h | 292.229 | 69.6115 | 0.097802 | 0.0166266 |
| Me-JA 6h | 274.637 | 67.5704 | 0.0912935 | 0.0187396 |
Source Transcript PGSC0003DMT400044098 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G32530.1 | +1 | 0.0 | 654 | 370/766 (48%) | cellulose synthase-like B3 | chr2:13809283-13813487 FORWARD LENGTH=755 |
