Probe CUST_44783_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44783_PI426222305 | JHI_St_60k_v1 | DMT400040632 | GGCTTTGGCTTATGAAAGATTGTTCCTTTGCATTAAGAATGAAACATTTTGTACCTACCA |
All Microarray Probes Designed to Gene DMG400015710
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44783_PI426222305 | JHI_St_60k_v1 | DMT400040632 | GGCTTTGGCTTATGAAAGATTGTTCCTTTGCATTAAGAATGAAACATTTTGTACCTACCA |
Microarray Signals from CUST_44783_PI426222305
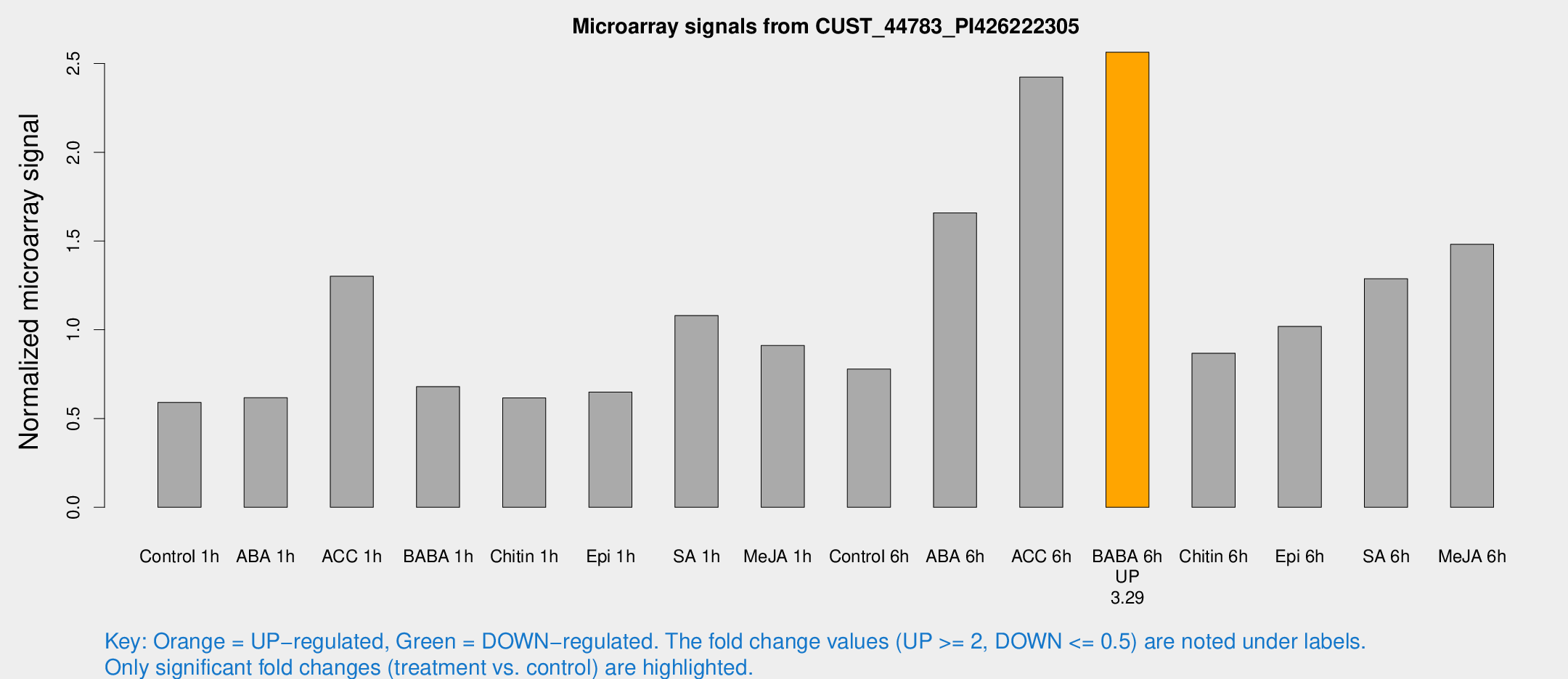
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 40.599 | 10.4994 | 0.590935 | 0.113174 |
| ABA 1h | 35.6133 | 3.88043 | 0.617792 | 0.0992541 |
| ACC 1h | 89.8265 | 16.8045 | 1.30151 | 0.175832 |
| BABA 1h | 47.1298 | 14.278 | 0.679773 | 0.171847 |
| Chitin 1h | 39.5011 | 12.939 | 0.6159 | 0.199237 |
| Epi 1h | 36.3967 | 4.03117 | 0.649357 | 0.0725947 |
| SA 1h | 75.5896 | 17.5207 | 1.08048 | 0.181397 |
| Me-JA 1h | 49.3506 | 8.14923 | 0.91141 | 0.155354 |
| Control 6h | 59.0669 | 20.1348 | 0.779035 | 0.279366 |
| ABA 6h | 132.704 | 53.3752 | 1.6587 | 0.586474 |
| ACC 6h | 183.569 | 27.2564 | 2.42335 | 0.150115 |
| BABA 6h | 198.001 | 51.4353 | 2.56347 | 0.630753 |
| Chitin 6h | 60.1056 | 5.24492 | 0.868134 | 0.0762472 |
| Epi 6h | 80.8319 | 23.6011 | 1.01908 | 0.226614 |
| SA 6h | 90.0197 | 23.2089 | 1.28798 | 0.262263 |
| Me-JA 6h | 95.4295 | 6.54854 | 1.48189 | 0.111929 |
Source Transcript PGSC0003DMT400040632 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G19460.1 | +1 | 2e-36 | 130 | 61/91 (67%) | UDP-Glycosyltransferase superfamily protein | chr4:10610422-10611972 REVERSE LENGTH=516 |
