Probe CUST_44719_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44719_PI426222305 | JHI_St_60k_v1 | DMT400045238 | GTTGGAGGAGATATGTTTGAAAGTGTTCCAGAAGGAGATGCTATTTTTATGAAGTTATTG |
All Microarray Probes Designed to Gene DMG400017552
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44704_PI426222305 | JHI_St_60k_v1 | DMT400045239 | CTGGACGGATAGTCATTGTGTGAAGTTATTGAAAAACTGTTATAAGTCTACACCAACAAA |
| CUST_44719_PI426222305 | JHI_St_60k_v1 | DMT400045238 | GTTGGAGGAGATATGTTTGAAAGTGTTCCAGAAGGAGATGCTATTTTTATGAAGTTATTG |
Microarray Signals from CUST_44719_PI426222305
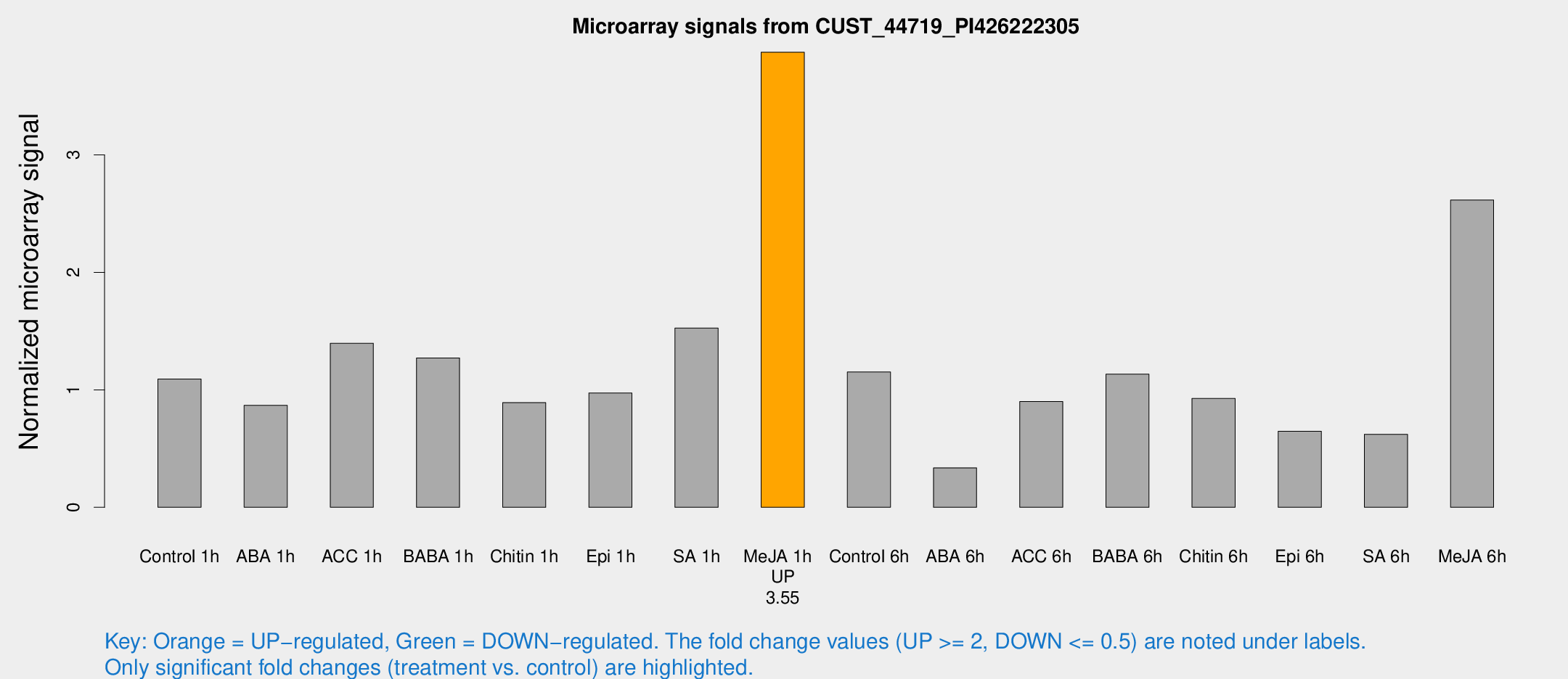
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 393.696 | 130.636 | 1.09145 | 0.286756 |
| ABA 1h | 256.189 | 40.7367 | 0.867852 | 0.0794606 |
| ACC 1h | 501.041 | 141.106 | 1.39535 | 0.334337 |
| BABA 1h | 404.562 | 48.0849 | 1.27125 | 0.0743113 |
| Chitin 1h | 280 | 67.5428 | 0.890997 | 0.210935 |
| Epi 1h | 287.917 | 67.5034 | 0.972896 | 0.191874 |
| SA 1h | 604.773 | 255.661 | 1.52622 | 0.74562 |
| Me-JA 1h | 1058.5 | 170.508 | 3.87361 | 0.358973 |
| Control 6h | 430.218 | 148.955 | 1.15177 | 0.355819 |
| ABA 6h | 135.692 | 53.8371 | 0.335697 | 0.165392 |
| ACC 6h | 352.004 | 74.0827 | 0.900623 | 0.134793 |
| BABA 6h | 433.622 | 87.607 | 1.13334 | 0.28646 |
| Chitin 6h | 337.946 | 77.6558 | 0.926155 | 0.20934 |
| Epi 6h | 246.851 | 47.0522 | 0.64711 | 0.170998 |
| SA 6h | 218.943 | 63.2415 | 0.621752 | 0.133824 |
| Me-JA 6h | 859.37 | 98.1657 | 2.61578 | 0.542902 |
Source Transcript PGSC0003DMT400045238 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G54160.1 | +1 | 5e-84 | 258 | 125/251 (50%) | O-methyltransferase 1 | chr5:21982075-21984167 FORWARD LENGTH=363 |
