Probe CUST_44704_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44704_PI426222305 | JHI_St_60k_v1 | DMT400045239 | CTGGACGGATAGTCATTGTGTGAAGTTATTGAAAAACTGTTATAAGTCTACACCAACAAA |
All Microarray Probes Designed to Gene DMG400017552
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44704_PI426222305 | JHI_St_60k_v1 | DMT400045239 | CTGGACGGATAGTCATTGTGTGAAGTTATTGAAAAACTGTTATAAGTCTACACCAACAAA |
| CUST_44719_PI426222305 | JHI_St_60k_v1 | DMT400045238 | GTTGGAGGAGATATGTTTGAAAGTGTTCCAGAAGGAGATGCTATTTTTATGAAGTTATTG |
Microarray Signals from CUST_44704_PI426222305
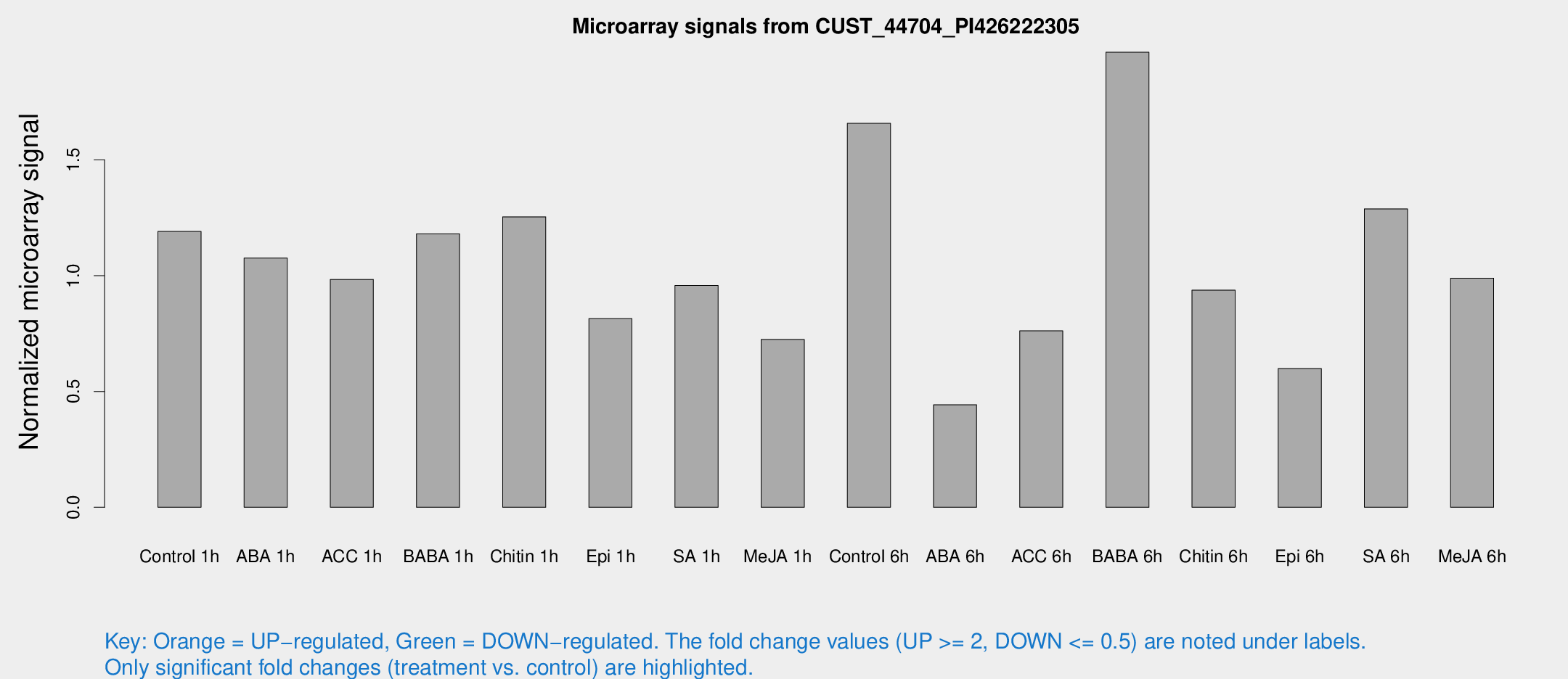
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18.0239 | 3.70285 | 1.19077 | 0.265 |
| ABA 1h | 17.7462 | 8.43846 | 1.07622 | 0.550466 |
| ACC 1h | 17.2718 | 6.6713 | 0.983297 | 0.378767 |
| BABA 1h | 17.435 | 4.14514 | 1.18057 | 0.292199 |
| Chitin 1h | 17.0035 | 3.68389 | 1.25384 | 0.290222 |
| Epi 1h | 12.5414 | 5.86928 | 0.814812 | 0.421621 |
| SA 1h | 15.4034 | 4.3538 | 0.957886 | 0.27031 |
| Me-JA 1h | 9.22427 | 3.7874 | 0.724572 | 0.345623 |
| Control 6h | 29.0032 | 11.7264 | 1.65758 | 0.675812 |
| ABA 6h | 6.8506 | 3.97401 | 0.442585 | 0.256425 |
| ACC 6h | 14.4665 | 4.73537 | 0.761888 | 0.292351 |
| BABA 6h | 34.1773 | 8.53222 | 1.96451 | 0.452871 |
| Chitin 6h | 15.6063 | 4.37923 | 0.937494 | 0.31003 |
| Epi 6h | 10.2782 | 4.42493 | 0.598921 | 0.269808 |
| SA 6h | 18.9656 | 4.21044 | 1.28844 | 0.354481 |
| Me-JA 6h | 17.6722 | 7.80139 | 0.988906 | 0.423728 |
Source Transcript PGSC0003DMT400045239 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G54160.1 | +3 | 1e-110 | 333 | 173/354 (49%) | O-methyltransferase 1 | chr5:21982075-21984167 FORWARD LENGTH=363 |
