Probe CUST_44514_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44514_PI426222305 | JHI_St_60k_v1 | DMT400043054 | CCCGGAAAGTGAAGTTCTTAAGAATAGAGTCTCCTCTAGTATCATATTGGCTAATGTTCT |
All Microarray Probes Designed to Gene DMG400016695
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44514_PI426222305 | JHI_St_60k_v1 | DMT400043054 | CCCGGAAAGTGAAGTTCTTAAGAATAGAGTCTCCTCTAGTATCATATTGGCTAATGTTCT |
Microarray Signals from CUST_44514_PI426222305
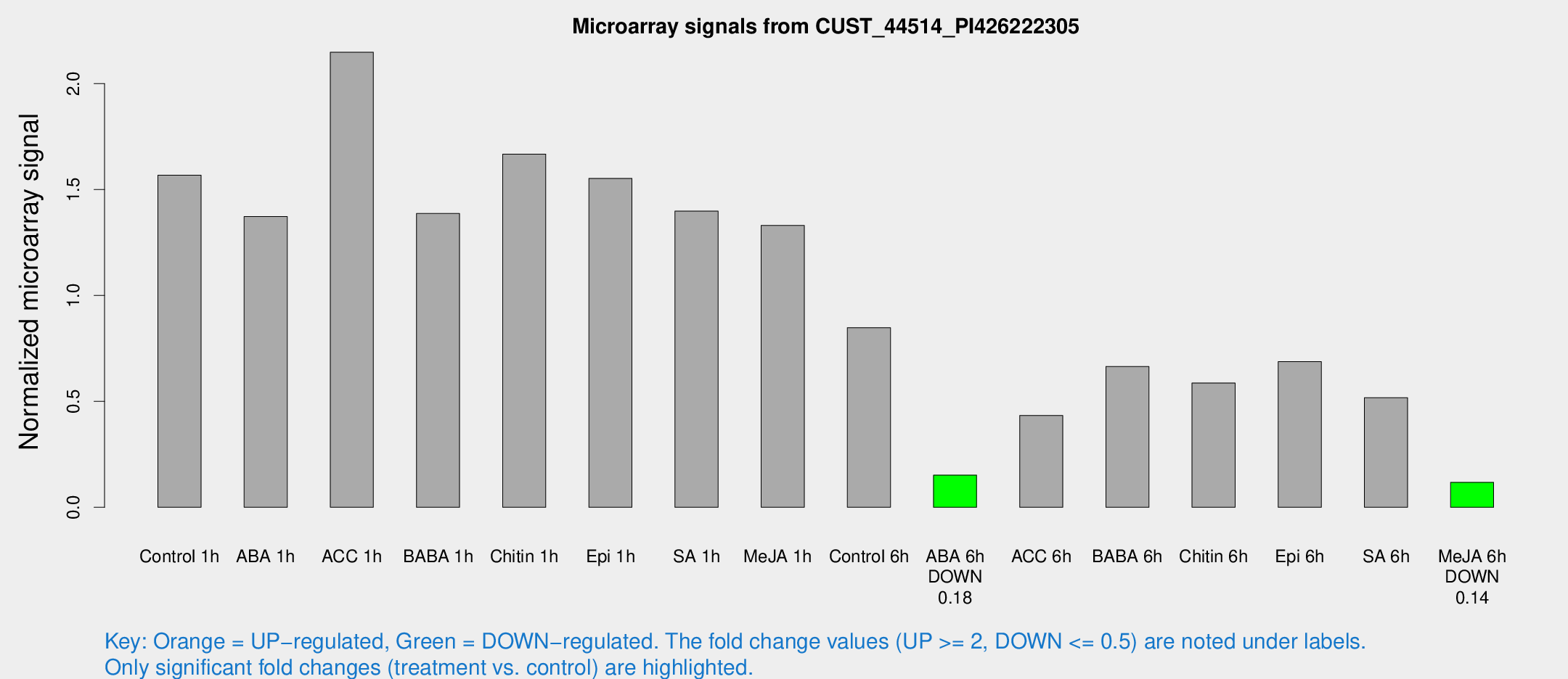
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 184324 | 28464.8 | 1.56816 | 0.188808 |
| ABA 1h | 143788 | 25920 | 1.37234 | 0.244614 |
| ACC 1h | 257002 | 33037.4 | 2.14843 | 0.422102 |
| BABA 1h | 155183 | 17998.7 | 1.38701 | 0.0800792 |
| Chitin 1h | 175755 | 28573 | 1.66752 | 0.278581 |
| Epi 1h | 155434 | 15849.2 | 1.55285 | 0.150999 |
| SA 1h | 166017 | 16317.2 | 1.3984 | 0.0807369 |
| Me-JA 1h | 125352 | 12291.7 | 1.33037 | 0.178206 |
| Control 6h | 105764 | 31010.4 | 0.848294 | 0.182826 |
| ABA 6h | 18780.2 | 2308.05 | 0.151945 | 0.0196137 |
| ACC 6h | 61745.4 | 18098.7 | 0.433534 | 0.0658362 |
| BABA 6h | 85924.6 | 8394.36 | 0.664809 | 0.0775296 |
| Chitin 6h | 72361.2 | 8360.19 | 0.586941 | 0.0708398 |
| Epi 6h | 94299.6 | 21340.7 | 0.688021 | 0.231938 |
| SA 6h | 63329.8 | 18086.1 | 0.517564 | 0.108243 |
| Me-JA 6h | 16291.4 | 6054.08 | 0.11784 | 0.0523433 |
Source Transcript PGSC0003DMT400043054 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G29930.1 | +2 | 3e-174 | 488 | 239/268 (89%) | chlorophyll A/B binding protein 1 | chr1:10478071-10478874 FORWARD LENGTH=267 |
