Probe CUST_44148_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44148_PI426222305 | JHI_St_60k_v1 | DMT400055239 | ATTAGTGGATTTGTTTGGAGTGGATATGGAATCAAGAAAGGTACCTTTGGCTTCTTTTTT |
All Microarray Probes Designed to Gene DMG400021444
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44148_PI426222305 | JHI_St_60k_v1 | DMT400055239 | ATTAGTGGATTTGTTTGGAGTGGATATGGAATCAAGAAAGGTACCTTTGGCTTCTTTTTT |
Microarray Signals from CUST_44148_PI426222305
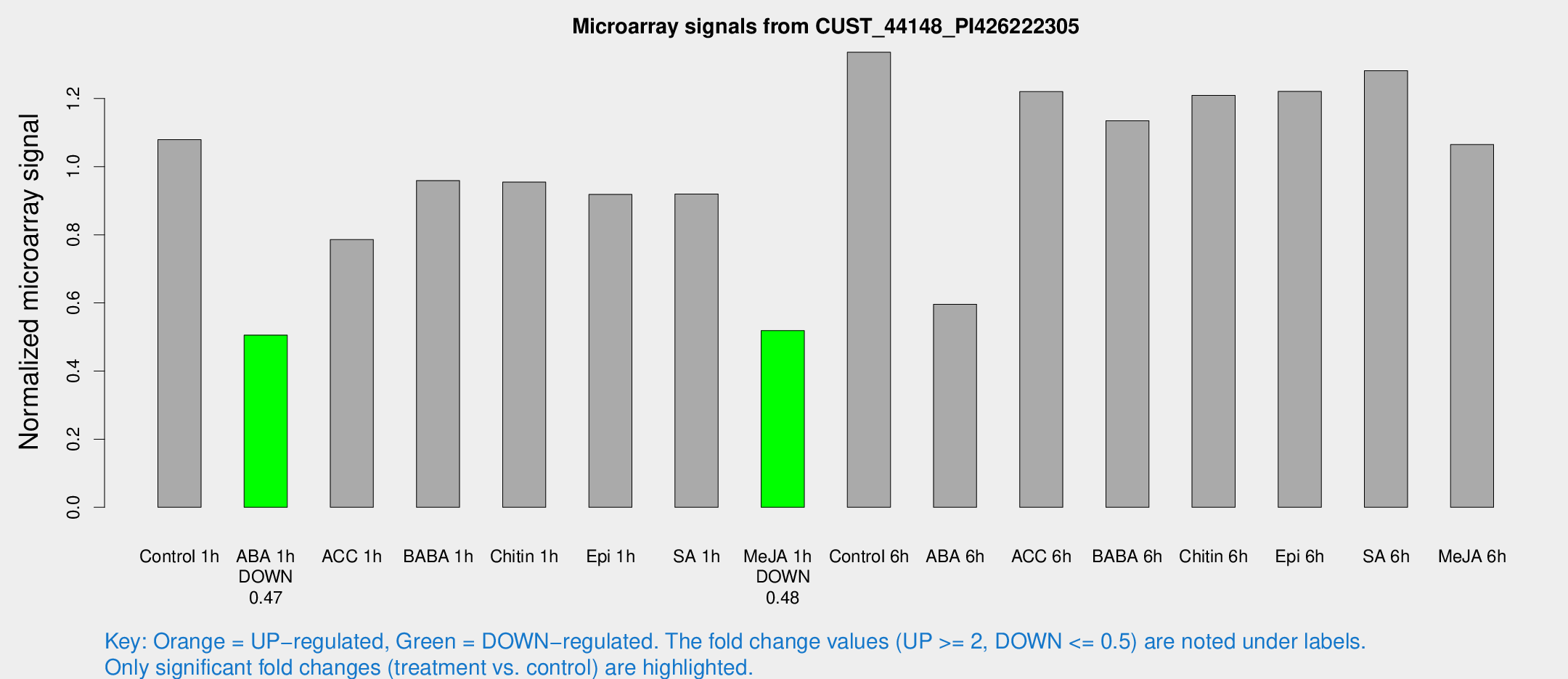
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 474.147 | 100.236 | 1.07928 | 0.150231 |
| ABA 1h | 188.707 | 11.3411 | 0.505501 | 0.03036 |
| ACC 1h | 372.438 | 97.7519 | 0.78594 | 0.197062 |
| BABA 1h | 401.615 | 67.2606 | 0.959244 | 0.081889 |
| Chitin 1h | 365.673 | 36.2771 | 0.954682 | 0.0557524 |
| Epi 1h | 336.408 | 20.6701 | 0.918438 | 0.0537378 |
| SA 1h | 411.829 | 75.7222 | 0.919868 | 0.153786 |
| Me-JA 1h | 179.805 | 14.6729 | 0.518833 | 0.0314878 |
| Control 6h | 586.862 | 109.857 | 1.33601 | 0.166409 |
| ABA 6h | 270.356 | 28.5788 | 0.595978 | 0.070156 |
| ACC 6h | 592.905 | 39.7621 | 1.22037 | 0.10932 |
| BABA 6h | 541.548 | 54.6655 | 1.13501 | 0.0790435 |
| Chitin 6h | 542.755 | 31.5591 | 1.20929 | 0.0703022 |
| Epi 6h | 589.769 | 71.9793 | 1.2208 | 0.0966214 |
| SA 6h | 558.653 | 110.465 | 1.28187 | 0.135143 |
| Me-JA 6h | 459.293 | 70.5715 | 1.0651 | 0.0971454 |
Source Transcript PGSC0003DMT400055239 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G37650.1 | +3 | 0.0 | 555 | 297/480 (62%) | GRAS family transcription factor | chr4:17691871-17693466 FORWARD LENGTH=531 |
