Probe CUST_44125_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44125_PI426222305 | JHI_St_60k_v1 | DMT400055176 | GTATGCTTAAATCTACGTCAATTAACGGATCTTTTCTTTTGTGGGTTCTTGGTCTAAATG |
All Microarray Probes Designed to Gene DMG400021414
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44125_PI426222305 | JHI_St_60k_v1 | DMT400055176 | GTATGCTTAAATCTACGTCAATTAACGGATCTTTTCTTTTGTGGGTTCTTGGTCTAAATG |
| CUST_44158_PI426222305 | JHI_St_60k_v1 | DMT400055175 | GTATGCTTAAATCTACGTCAATTAACGGATCTTTTCTTTTGTGGGTTCTTGGTCTAAATG |
Microarray Signals from CUST_44125_PI426222305
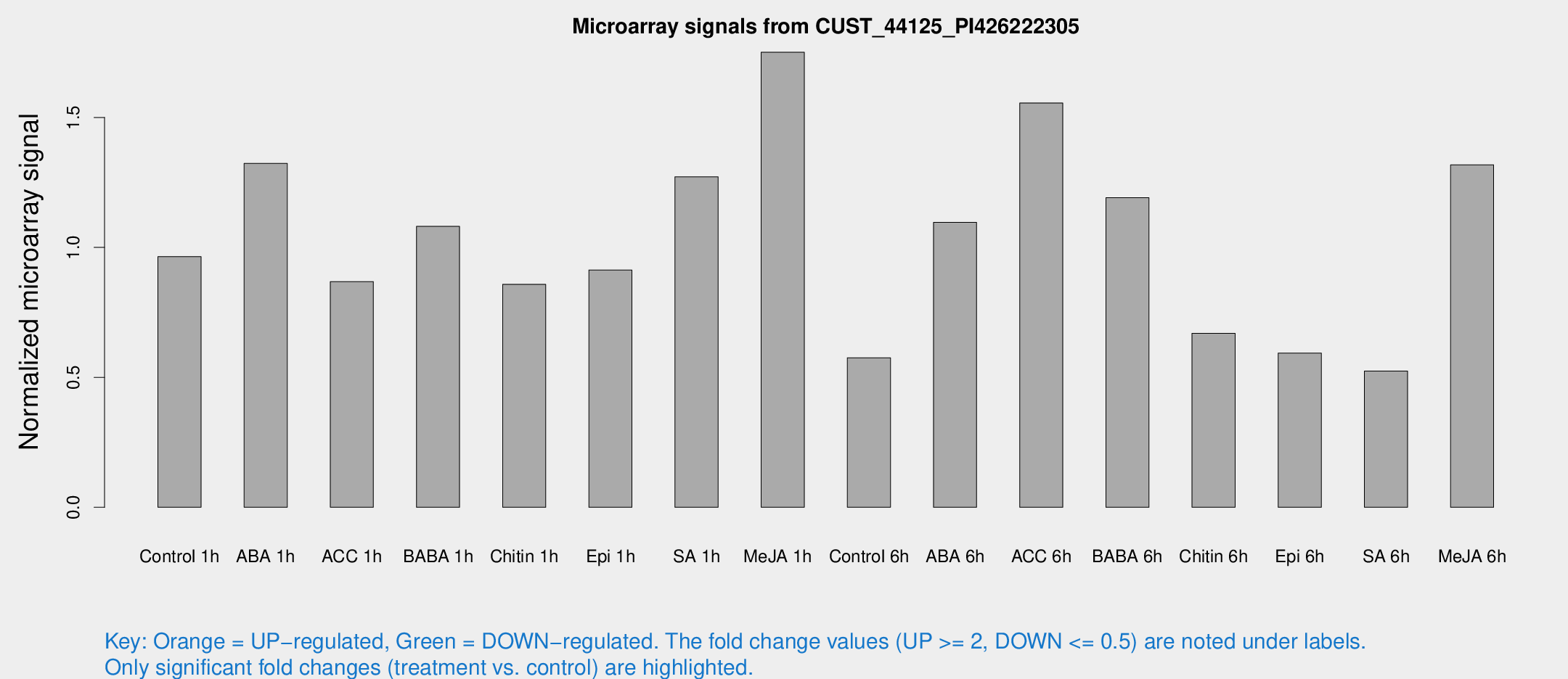
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 302.129 | 25.7413 | 0.964675 | 0.0570561 |
| ABA 1h | 377.881 | 66.9809 | 1.32354 | 0.192005 |
| ACC 1h | 306.891 | 86.8351 | 0.868372 | 0.240329 |
| BABA 1h | 334.849 | 56.7484 | 1.08092 | 0.0904984 |
| Chitin 1h | 241.53 | 19.0548 | 0.857981 | 0.0513668 |
| Epi 1h | 249.409 | 29.5884 | 0.912928 | 0.0924848 |
| SA 1h | 410.985 | 45.4791 | 1.27148 | 0.161059 |
| Me-JA 1h | 449.138 | 46.2821 | 1.75108 | 0.102191 |
| Control 6h | 191.313 | 45.1168 | 0.575558 | 0.0983593 |
| ABA 6h | 377.513 | 70.2418 | 1.09643 | 0.177689 |
| ACC 6h | 562.561 | 66.8988 | 1.55572 | 0.35467 |
| BABA 6h | 439.093 | 108.198 | 1.19132 | 0.265118 |
| Chitin 6h | 225.822 | 31.8651 | 0.669459 | 0.092014 |
| Epi 6h | 208.489 | 12.9354 | 0.593344 | 0.0367037 |
| SA 6h | 183.929 | 56.5693 | 0.524507 | 0.150367 |
| Me-JA 6h | 408.712 | 23.9258 | 1.31782 | 0.120622 |
Source Transcript PGSC0003DMT400055176 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G22910.1 | +1 | 0.0 | 756 | 375/538 (70%) | N-acetyl-l-glutamate synthase 1 | chr2:9749988-9752737 FORWARD LENGTH=609 |
