Probe CUST_43712_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43712_PI426222305 | JHI_St_60k_v1 | DMT400004892 | TCTACCTTACCACTTCTTGCTCAGGCAAAACCAAGATTAGCTAAGGTCATGTATCTTTAA |
All Microarray Probes Designed to Gene DMG400001944
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43691_PI426222305 | JHI_St_60k_v1 | DMT400004893 | CTTCTTGCTCAGGCAAAACCAAGATTAGCTAAGGTCATGTATCTTTAAGCTACAATATTT |
| CUST_43712_PI426222305 | JHI_St_60k_v1 | DMT400004892 | TCTACCTTACCACTTCTTGCTCAGGCAAAACCAAGATTAGCTAAGGTCATGTATCTTTAA |
Microarray Signals from CUST_43712_PI426222305
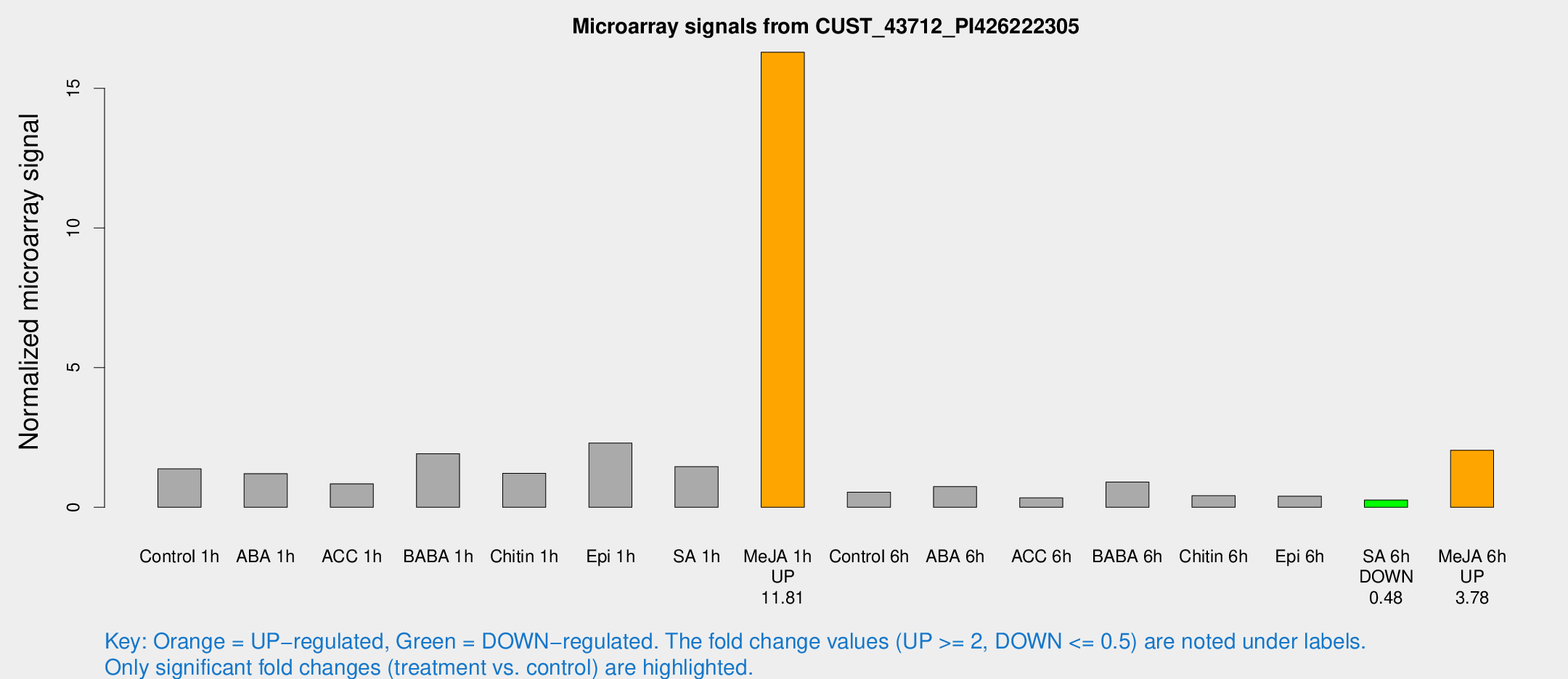
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 35.4057 | 9.22566 | 1.37946 | 0.485678 |
| ABA 1h | 45.9961 | 29.731 | 1.20587 | 1.67059 |
| ACC 1h | 21.1245 | 4.54992 | 0.840486 | 0.177675 |
| BABA 1h | 51.5082 | 17.4008 | 1.91933 | 0.738071 |
| Chitin 1h | 30.5017 | 10.8505 | 1.21771 | 0.759935 |
| Epi 1h | 51.0611 | 13.2782 | 2.29888 | 0.755389 |
| SA 1h | 37.5223 | 10.0022 | 1.45927 | 0.470189 |
| Me-JA 1h | 316.856 | 41.8616 | 16.2894 | 1.24746 |
| Control 6h | 13.3301 | 3.4831 | 0.540878 | 0.159598 |
| ABA 6h | 18.5245 | 3.64453 | 0.741533 | 0.146525 |
| ACC 6h | 11.3576 | 5.60173 | 0.337476 | 0.160247 |
| BABA 6h | 26.3168 | 7.27828 | 0.90244 | 0.358803 |
| Chitin 6h | 13.094 | 6.55675 | 0.416095 | 0.246668 |
| Epi 6h | 10.6314 | 3.87801 | 0.395027 | 0.155466 |
| SA 6h | 6.07038 | 3.51803 | 0.261772 | 0.15168 |
| Me-JA 6h | 49.5702 | 9.55747 | 2.04359 | 0.330399 |
Source Transcript PGSC0003DMT400004892 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G05260.1 | +1 | 1e-85 | 270 | 154/289 (53%) | cytochrome p450 79a2 | chr5:1559778-1561765 REVERSE LENGTH=523 |
