Probe CUST_43691_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43691_PI426222305 | JHI_St_60k_v1 | DMT400004893 | CTTCTTGCTCAGGCAAAACCAAGATTAGCTAAGGTCATGTATCTTTAAGCTACAATATTT |
All Microarray Probes Designed to Gene DMG400001944
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43691_PI426222305 | JHI_St_60k_v1 | DMT400004893 | CTTCTTGCTCAGGCAAAACCAAGATTAGCTAAGGTCATGTATCTTTAAGCTACAATATTT |
| CUST_43712_PI426222305 | JHI_St_60k_v1 | DMT400004892 | TCTACCTTACCACTTCTTGCTCAGGCAAAACCAAGATTAGCTAAGGTCATGTATCTTTAA |
Microarray Signals from CUST_43691_PI426222305
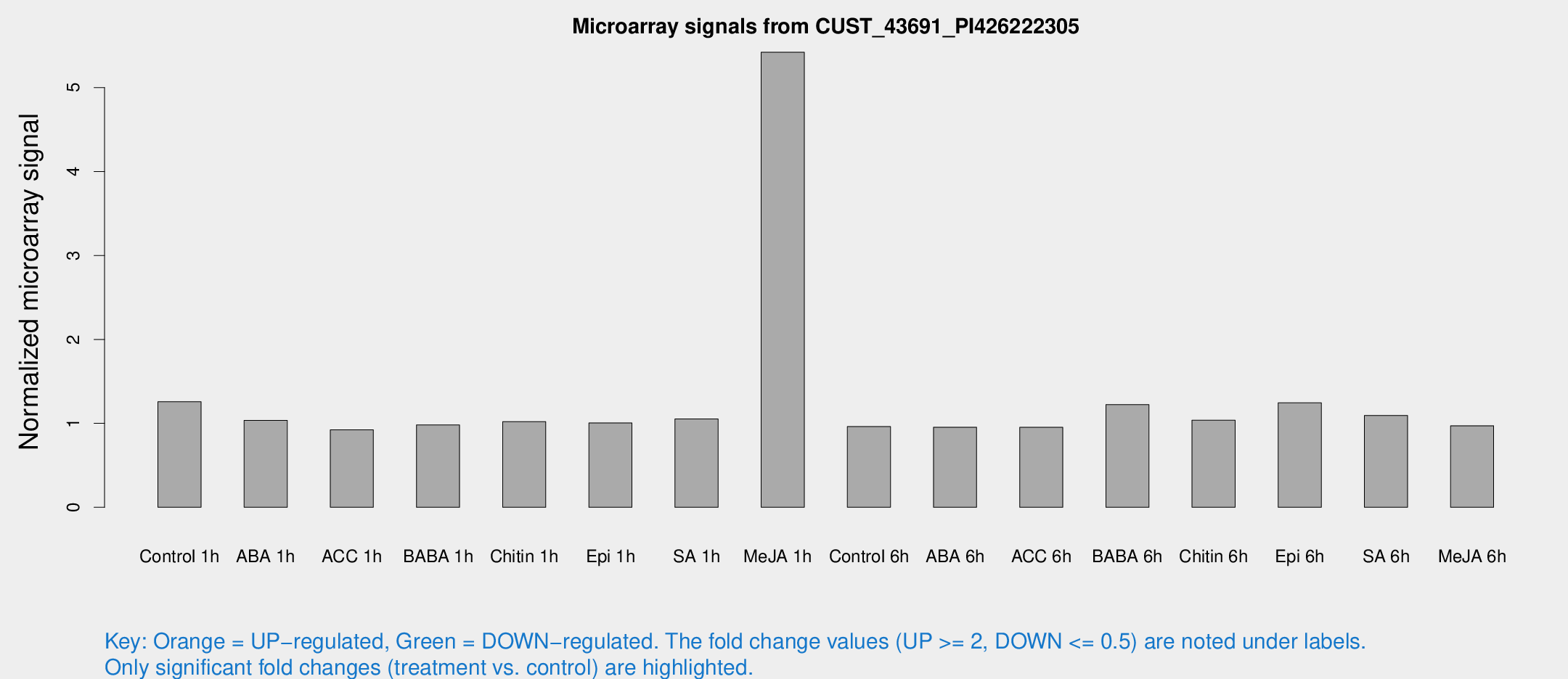
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8.63381 | 3.45992 | 1.25738 | 0.540146 |
| ABA 1h | 6.12497 | 3.27661 | 1.03573 | 0.560125 |
| ACC 1h | 6.30372 | 3.65305 | 0.922512 | 0.534244 |
| BABA 1h | 6.29815 | 3.65394 | 0.981305 | 0.568494 |
| Chitin 1h | 6.14157 | 3.45464 | 1.02063 | 0.57107 |
| Epi 1h | 5.79001 | 3.35804 | 1.0045 | 0.581713 |
| SA 1h | 7.48459 | 3.34001 | 1.052 | 0.515644 |
| Me-JA 1h | 30.8101 | 6.25864 | 5.42119 | 1.16368 |
| Control 6h | 6.41585 | 3.71988 | 0.961535 | 0.556834 |
| ABA 6h | 6.75063 | 3.91198 | 0.954193 | 0.552534 |
| ACC 6h | 7.40725 | 4.37972 | 0.9526 | 0.552236 |
| BABA 6h | 10.2765 | 4.20281 | 1.22332 | 0.593106 |
| Chitin 6h | 7.35153 | 4.26154 | 1.03745 | 0.600724 |
| Epi 6h | 9.74457 | 4.29551 | 1.24453 | 0.600906 |
| SA 6h | 7.2119 | 3.98941 | 1.0945 | 0.606363 |
| Me-JA 6h | 6.44501 | 3.74288 | 0.971131 | 0.56272 |
Source Transcript PGSC0003DMT400004893 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G05260.1 | +3 | 9e-121 | 301 | 174/347 (50%) | cytochrome p450 79a2 | chr5:1559778-1561765 REVERSE LENGTH=523 |
