Probe CUST_43565_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43565_PI426222305 | JHI_St_60k_v1 | DMT400064750 | GTTTATATTGGAGAAGCTTTTGAGCTTCCATGTAATGTAGCACTTGAACCAAGTAATAAC |
All Microarray Probes Designed to Gene DMG402025150
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43564_PI426222305 | JHI_St_60k_v1 | DMT400064751 | GTTTATATTGGAGAAGCTTTTGAGCTTCCATGTAATGTAGCACTTGAACCAAGTAATAAC |
| CUST_43565_PI426222305 | JHI_St_60k_v1 | DMT400064750 | GTTTATATTGGAGAAGCTTTTGAGCTTCCATGTAATGTAGCACTTGAACCAAGTAATAAC |
Microarray Signals from CUST_43565_PI426222305
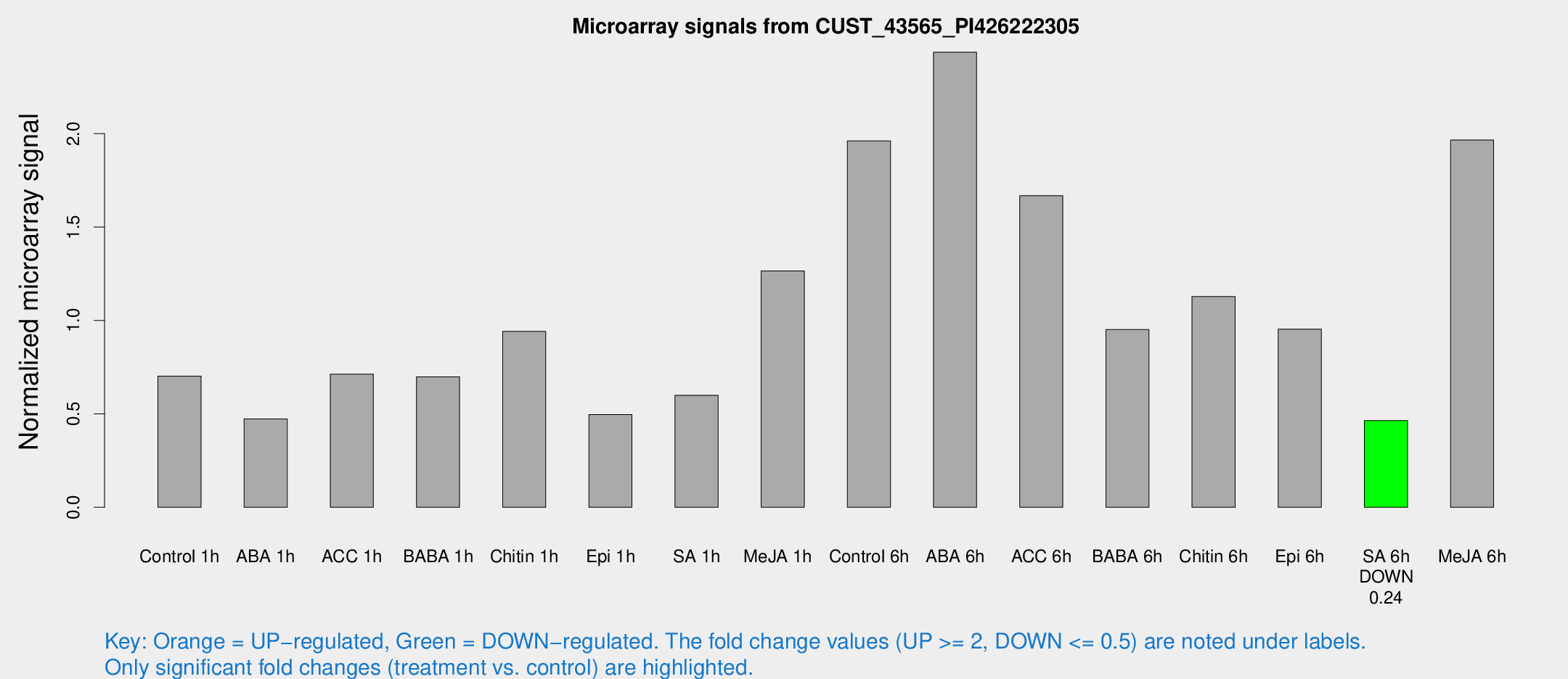
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18.8211 | 7.53614 | 0.701981 | 0.356351 |
| ABA 1h | 10.3049 | 3.47365 | 0.473469 | 0.187939 |
| ACC 1h | 19.1655 | 6.5395 | 0.712895 | 0.239666 |
| BABA 1h | 17.5334 | 5.9582 | 0.698395 | 0.213661 |
| Chitin 1h | 19.7784 | 3.73802 | 0.941934 | 0.182176 |
| Epi 1h | 10.384 | 3.53132 | 0.495938 | 0.192997 |
| SA 1h | 14.2498 | 3.59416 | 0.599768 | 0.154058 |
| Me-JA 1h | 23.815 | 3.93016 | 1.26495 | 0.216118 |
| Control 6h | 46.3103 | 8.04486 | 1.96114 | 0.262415 |
| ABA 6h | 60.3468 | 7.58382 | 2.43604 | 0.218881 |
| ACC 6h | 43.9106 | 5.18493 | 1.66857 | 0.19481 |
| BABA 6h | 25.8626 | 5.91713 | 0.951671 | 0.202312 |
| Chitin 6h | 28.2276 | 4.70204 | 1.12801 | 0.195531 |
| Epi 6h | 30.149 | 10.8588 | 0.953686 | 0.678222 |
| SA 6h | 10.5758 | 4.13025 | 0.464181 | 0.182864 |
| Me-JA 6h | 45.6134 | 5.58144 | 1.96614 | 0.340145 |
Source Transcript PGSC0003DMT400064750 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G29180.1 | +1 | 1e-159 | 481 | 265/478 (55%) | Protein of unknown function (DUF1336) | chr3:11149073-11151322 FORWARD LENGTH=513 |
