Probe CUST_43522_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43522_PI426222305 | JHI_St_60k_v1 | DMT400065369 | ATCCCATGCCCATGGGCTACTTTACTGAAACAGTGATGGGAGCTTACAACTGCTTTGATT |
All Microarray Probes Designed to Gene DMG400025409
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43522_PI426222305 | JHI_St_60k_v1 | DMT400065369 | ATCCCATGCCCATGGGCTACTTTACTGAAACAGTGATGGGAGCTTACAACTGCTTTGATT |
| CUST_43518_PI426222305 | JHI_St_60k_v1 | DMT400065370 | TGCTGGGATTACAAAGTATATTGTACTTCCCCTCGCACATTTGAACGTTTATATATATGT |
Microarray Signals from CUST_43522_PI426222305
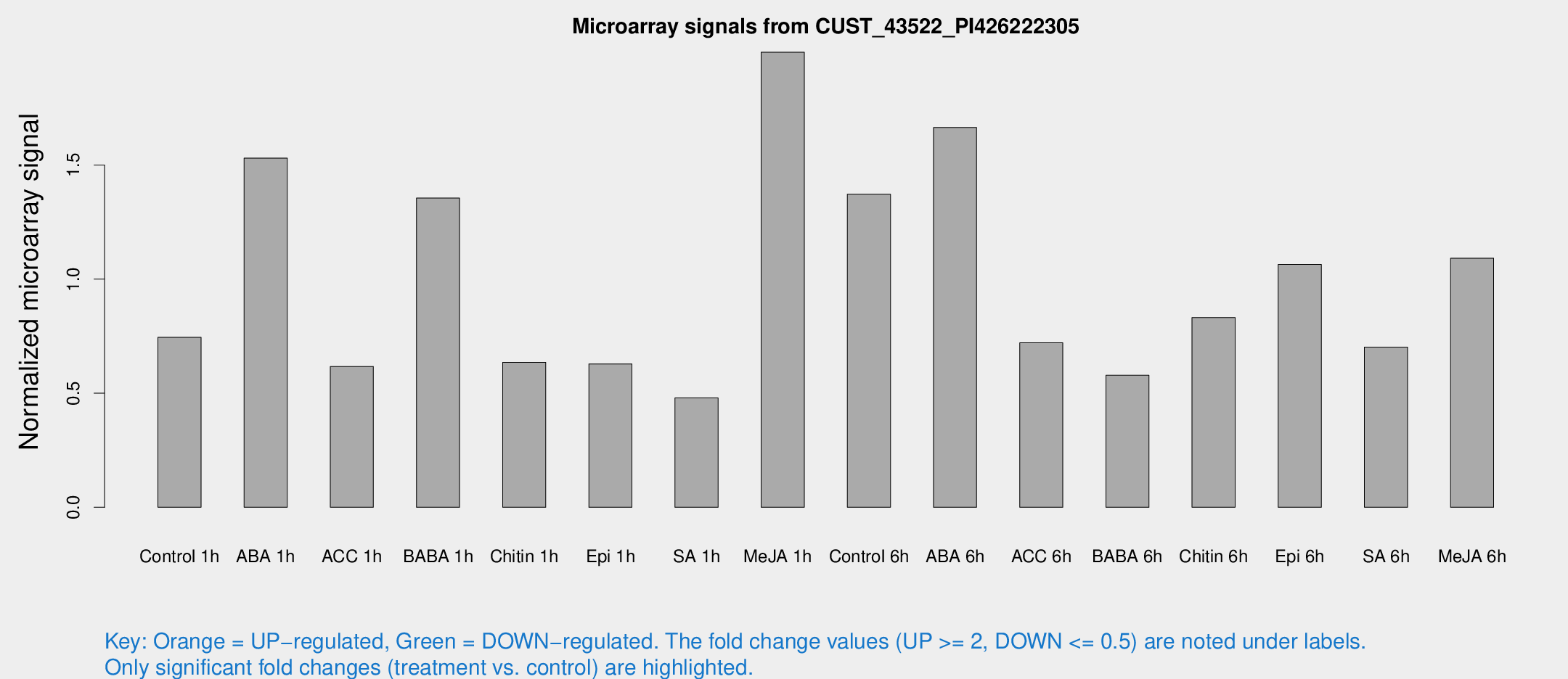
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.5003 | 3.17242 | 0.745046 | 0.302489 |
| ABA 1h | 18.8126 | 8.14838 | 1.53067 | 0.732943 |
| ACC 1h | 7.92784 | 3.14308 | 0.61735 | 0.293589 |
| BABA 1h | 15.0184 | 3.14803 | 1.35541 | 0.284032 |
| Chitin 1h | 6.91821 | 2.90887 | 0.635762 | 0.304423 |
| Epi 1h | 6.4338 | 2.89176 | 0.628578 | 0.301398 |
| SA 1h | 5.66803 | 2.89454 | 0.479619 | 0.246388 |
| Me-JA 1h | 18.8962 | 3.14084 | 1.99485 | 0.335104 |
| Control 6h | 16.1148 | 3.13086 | 1.37276 | 0.281208 |
| ABA 6h | 22.1336 | 5.65048 | 1.66495 | 0.461007 |
| ACC 6h | 10.2756 | 3.67415 | 0.721198 | 0.293195 |
| BABA 6h | 8.17423 | 3.39945 | 0.578887 | 0.280858 |
| Chitin 6h | 11.4122 | 3.75001 | 0.832168 | 0.331744 |
| Epi 6h | 14.7469 | 3.75941 | 1.06476 | 0.344517 |
| SA 6h | 9.14683 | 3.55856 | 0.702476 | 0.326797 |
| Me-JA 6h | 16.0229 | 7.71178 | 1.09197 | 0.619609 |
Source Transcript PGSC0003DMT400065369 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
