Probe CUST_43413_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43413_PI426222305 | JHI_St_60k_v1 | DMT400033012 | CAGATAATAATGGGAGGTCACCGAGGAAAACGCCTGGTAATCGTCTCTGTTAATCTTCAT |
All Microarray Probes Designed to Gene DMG400012676
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43395_PI426222305 | JHI_St_60k_v1 | DMT400033013 | GCCCAAACTGCCACATTACAAATAGCCAATCAGCTGTTATTTTTATCTCCAAGTACAAAA |
| CUST_43413_PI426222305 | JHI_St_60k_v1 | DMT400033012 | CAGATAATAATGGGAGGTCACCGAGGAAAACGCCTGGTAATCGTCTCTGTTAATCTTCAT |
Microarray Signals from CUST_43413_PI426222305
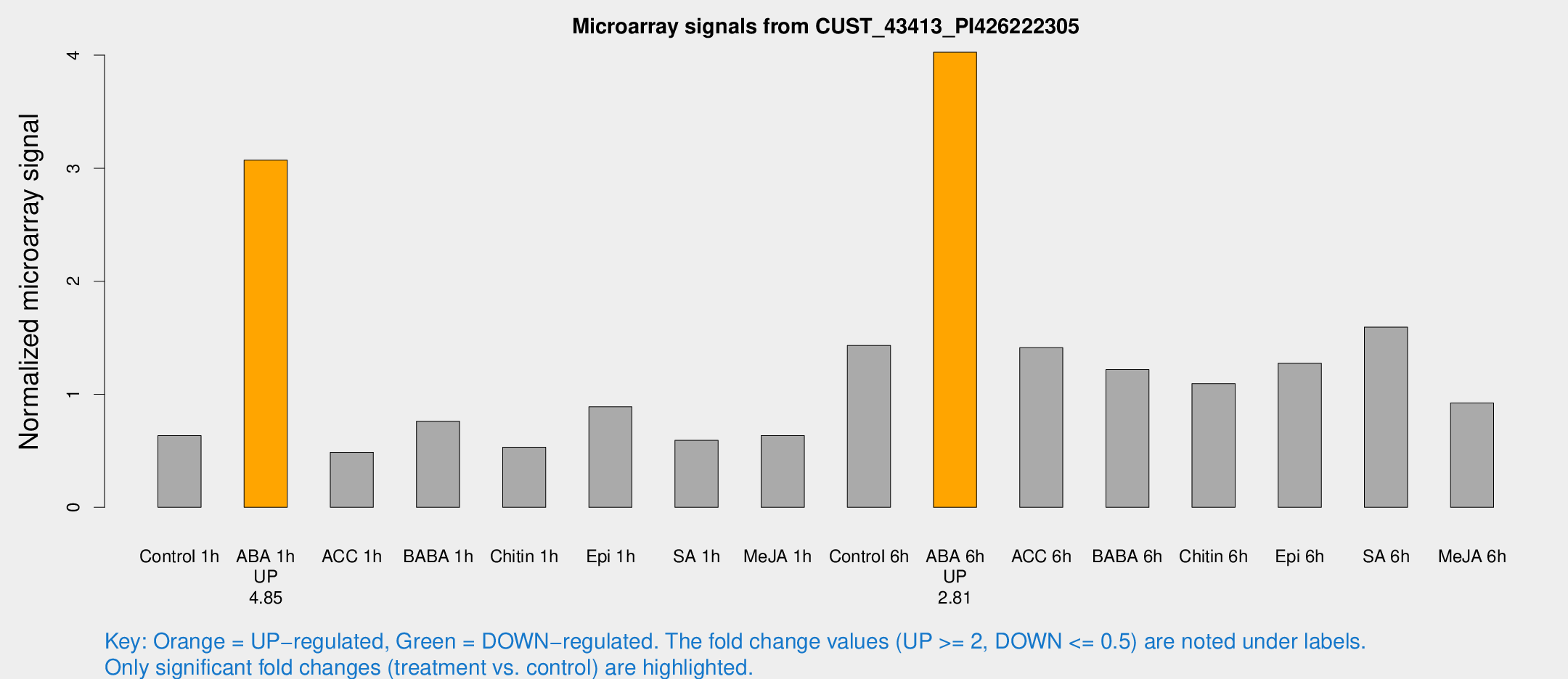
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 48.998 | 4.30379 | 0.633789 | 0.0556996 |
| ABA 1h | 211.873 | 20.4242 | 3.07098 | 0.225637 |
| ACC 1h | 39.9812 | 6.86511 | 0.486973 | 0.0619905 |
| BABA 1h | 57.6167 | 7.3277 | 0.761207 | 0.0642222 |
| Chitin 1h | 37.3249 | 4.02846 | 0.531601 | 0.0571575 |
| Epi 1h | 61.0317 | 9.52248 | 0.888949 | 0.143509 |
| SA 1h | 48.1581 | 7.571 | 0.593182 | 0.0911463 |
| Me-JA 1h | 40.2911 | 4.10234 | 0.634251 | 0.111473 |
| Control 6h | 115.293 | 24.0828 | 1.43107 | 0.283855 |
| ABA 6h | 335.737 | 41.3022 | 4.02642 | 0.281231 |
| ACC 6h | 134.702 | 37.7071 | 1.4123 | 0.220755 |
| BABA 6h | 108.74 | 20.4241 | 1.21761 | 0.209766 |
| Chitin 6h | 92.9035 | 15.7794 | 1.09506 | 0.231268 |
| Epi 6h | 112.986 | 15.0879 | 1.27516 | 0.149495 |
| SA 6h | 121.824 | 7.8585 | 1.59458 | 0.189974 |
| Me-JA 6h | 77.5842 | 22.1727 | 0.923782 | 0.207772 |
Source Transcript PGSC0003DMT400033012 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G04560.1 | +1 | 9e-17 | 74 | 64/117 (55%) | AWPM-19-like family protein | chr1:1245070-1245888 FORWARD LENGTH=186 |
