Probe CUST_43395_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43395_PI426222305 | JHI_St_60k_v1 | DMT400033013 | GCCCAAACTGCCACATTACAAATAGCCAATCAGCTGTTATTTTTATCTCCAAGTACAAAA |
All Microarray Probes Designed to Gene DMG400012676
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43395_PI426222305 | JHI_St_60k_v1 | DMT400033013 | GCCCAAACTGCCACATTACAAATAGCCAATCAGCTGTTATTTTTATCTCCAAGTACAAAA |
| CUST_43413_PI426222305 | JHI_St_60k_v1 | DMT400033012 | CAGATAATAATGGGAGGTCACCGAGGAAAACGCCTGGTAATCGTCTCTGTTAATCTTCAT |
Microarray Signals from CUST_43395_PI426222305
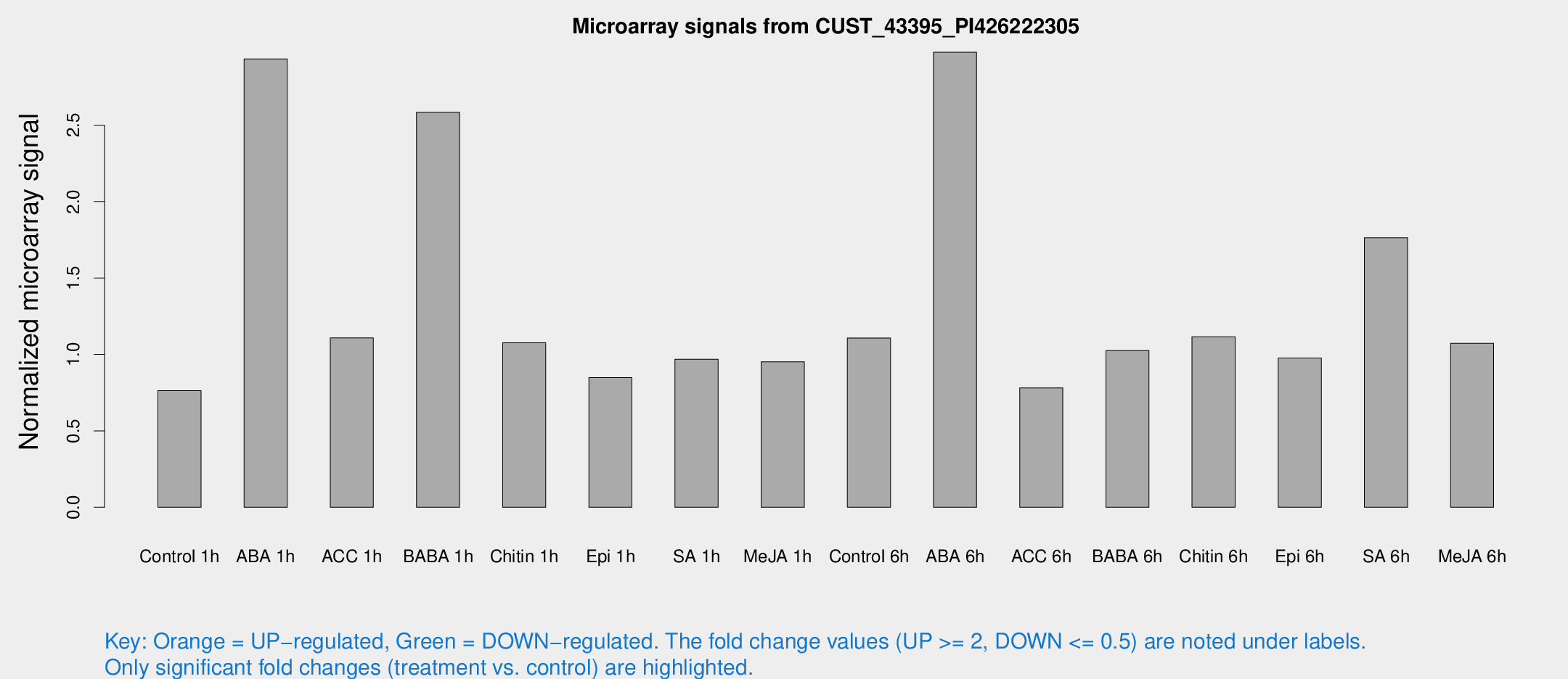
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.13304 | 3.56936 | 0.762569 | 0.442098 |
| ABA 1h | 21.7818 | 4.80949 | 2.93232 | 0.542162 |
| ACC 1h | 9.17017 | 4.62326 | 1.10779 | 0.538339 |
| BABA 1h | 23.9652 | 10.5062 | 2.58448 | 1.19241 |
| Chitin 1h | 8.31866 | 3.58249 | 1.07607 | 0.522741 |
| Epi 1h | 5.91107 | 3.4413 | 0.848544 | 0.491793 |
| SA 1h | 8.17631 | 3.41369 | 0.968591 | 0.42719 |
| Me-JA 1h | 6.28082 | 3.67123 | 0.951649 | 0.551017 |
| Control 6h | 9.62829 | 4.10292 | 1.10768 | 0.541079 |
| ABA 6h | 25.4468 | 4.1322 | 2.97675 | 0.486339 |
| ACC 6h | 7.35805 | 4.37382 | 0.781431 | 0.453063 |
| BABA 6h | 9.69009 | 4.17103 | 1.02479 | 0.496029 |
| Chitin 6h | 10.1942 | 4.16929 | 1.11601 | 0.533335 |
| Epi 6h | 8.88011 | 4.37049 | 0.97632 | 0.474225 |
| SA 6h | 14.6726 | 4.0527 | 1.76274 | 0.543011 |
| Me-JA 6h | 9.50727 | 3.7582 | 1.07249 | 0.513058 |
Source Transcript PGSC0003DMT400033013 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G04560.1 | +3 | 7e-26 | 107 | 81/148 (55%) | AWPM-19-like family protein | chr1:1245070-1245888 FORWARD LENGTH=186 |
