Probe CUST_42597_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42597_PI426222305 | JHI_St_60k_v1 | DMT400026850 | GGTGATGCTGTCCTTGATGCCTCAATGTAAAATTGATATTTCTTCAACAAGTACATAACA |
All Microarray Probes Designed to Gene DMG400010362
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42597_PI426222305 | JHI_St_60k_v1 | DMT400026850 | GGTGATGCTGTCCTTGATGCCTCAATGTAAAATTGATATTTCTTCAACAAGTACATAACA |
| CUST_42594_PI426222305 | JHI_St_60k_v1 | DMT400026852 | GGATTGCCTTCAGCAAGAAATCTAAATGTGGAACTTGATGATGATTTCCTTAATGTATCA |
Microarray Signals from CUST_42597_PI426222305
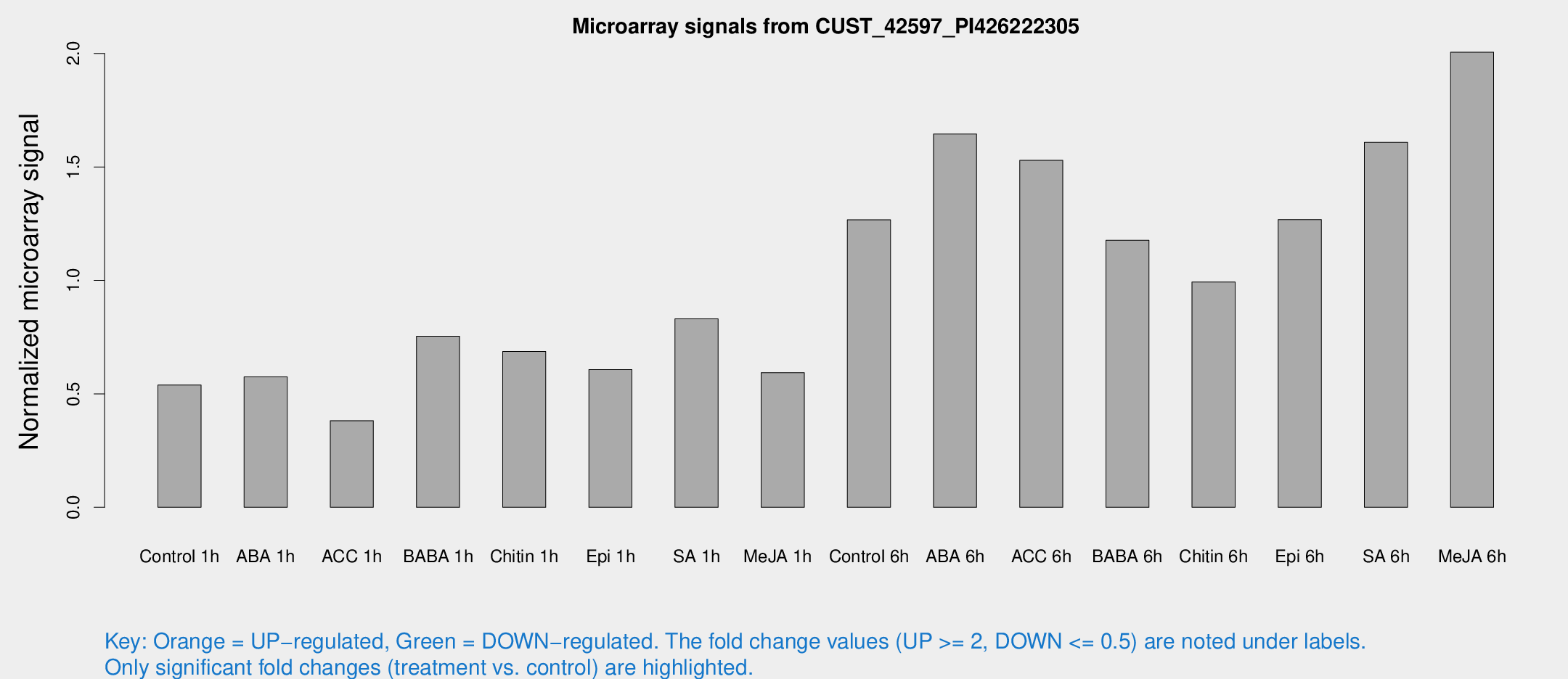
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 14230.1 | 1396.83 | 0.539032 | 0.0382575 |
| ABA 1h | 14052.1 | 2966.33 | 0.574808 | 0.113923 |
| ACC 1h | 11991 | 4094.38 | 0.381828 | 0.147464 |
| BABA 1h | 20812.6 | 5481.86 | 0.753655 | 0.168294 |
| Chitin 1h | 16696.9 | 3059.56 | 0.686454 | 0.109176 |
| Epi 1h | 14144.4 | 2399.71 | 0.607243 | 0.104225 |
| SA 1h | 23285.7 | 4481.43 | 0.830972 | 0.128066 |
| Me-JA 1h | 12961.1 | 1993.66 | 0.593192 | 0.038871 |
| Control 6h | 33872 | 4553.05 | 1.26686 | 0.0808436 |
| ABA 6h | 47815.2 | 9369.7 | 1.645 | 0.280259 |
| ACC 6h | 45874.1 | 2648.63 | 1.52892 | 0.204861 |
| BABA 6h | 34532.5 | 1998.92 | 1.17645 | 0.0679226 |
| Chitin 6h | 27658.2 | 1597.77 | 0.992882 | 0.0573243 |
| Epi 6h | 37816.6 | 3838.62 | 1.26811 | 0.0732144 |
| SA 6h | 44111.9 | 9803.13 | 1.60808 | 0.221889 |
| Me-JA 6h | 52521.9 | 3803.04 | 2.0056 | 0.115794 |
Source Transcript PGSC0003DMT400026850 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G02300.1 | +1 | 5e-163 | 472 | 228/398 (57%) | Cysteine proteinases superfamily protein | chr1:453288-455376 FORWARD LENGTH=379 |
