Probe CUST_42440_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42440_PI426222305 | JHI_St_60k_v1 | DMT400059171 | AAGGAATCCCCGTTCTCCGATGACATTCAGGGTTCTGATGCTACAGGAAATGATGTACAA |
All Microarray Probes Designed to Gene DMG400022983
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42459_PI426222305 | JHI_St_60k_v1 | DMT400059170 | AAGGAATCCCCGTTCTCCGATGACATTCAGGGTTCTGATGCTACAGGAAATGATGTACAA |
| CUST_42440_PI426222305 | JHI_St_60k_v1 | DMT400059171 | AAGGAATCCCCGTTCTCCGATGACATTCAGGGTTCTGATGCTACAGGAAATGATGTACAA |
Microarray Signals from CUST_42440_PI426222305
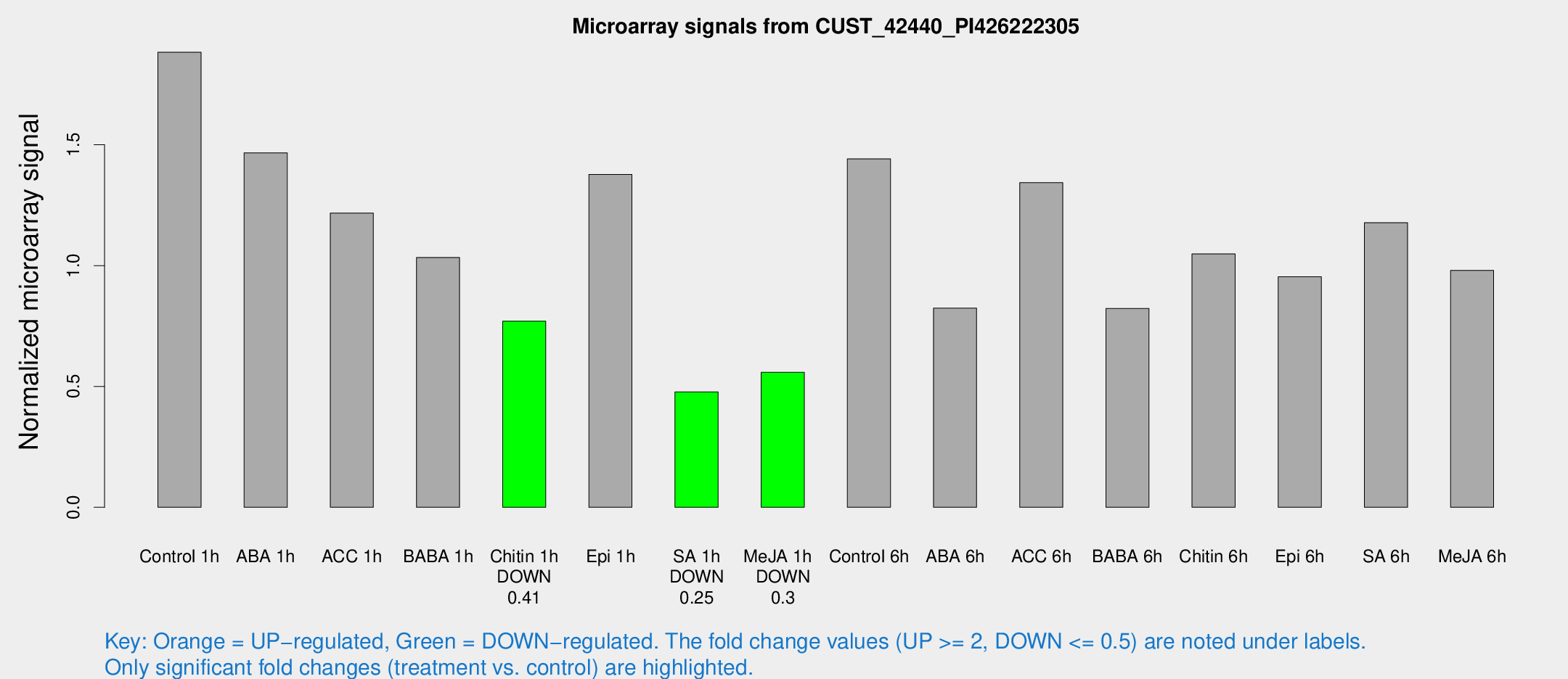
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3597.77 | 830.05 | 1.88284 | 0.3594 |
| ABA 1h | 2669.88 | 1027.23 | 1.46675 | 0.593417 |
| ACC 1h | 2531.83 | 743.809 | 1.21736 | 0.374312 |
| BABA 1h | 2008.11 | 571.327 | 1.03355 | 0.27832 |
| Chitin 1h | 1275.92 | 182.409 | 0.770179 | 0.0786745 |
| Epi 1h | 2148.31 | 124.184 | 1.37745 | 0.0795528 |
| SA 1h | 914.102 | 160.361 | 0.477174 | 0.0654424 |
| Me-JA 1h | 843.391 | 133.149 | 0.558468 | 0.0480106 |
| Control 6h | 2928.61 | 1004.95 | 1.44165 | 0.432849 |
| ABA 6h | 1578.74 | 91.3104 | 0.823962 | 0.0729611 |
| ACC 6h | 2847.9 | 430.843 | 1.34333 | 0.191122 |
| BABA 6h | 1782.14 | 478.53 | 0.82285 | 0.21158 |
| Chitin 6h | 2087.44 | 424.12 | 1.04822 | 0.172712 |
| Epi 6h | 1974.81 | 278.548 | 0.953933 | 0.114534 |
| SA 6h | 2219.91 | 529.711 | 1.17743 | 0.180474 |
| Me-JA 6h | 1801.13 | 286.836 | 0.980272 | 0.0753561 |
Source Transcript PGSC0003DMT400059171 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G57180.2 | +3 | 2e-63 | 219 | 189/436 (43%) | chloroplast import apparatus 2 | chr5:23168393-23170763 FORWARD LENGTH=435 |
