Probe CUST_4203_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4203_PI426222305 | JHI_St_60k_v1 | DMT400061678 | GCACTATCGATATCCAAACAAGGAAAACAATAAGCACATCAATGTAAAGGAGATACTAGA |
All Microarray Probes Designed to Gene DMG400024003
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4203_PI426222305 | JHI_St_60k_v1 | DMT400061678 | GCACTATCGATATCCAAACAAGGAAAACAATAAGCACATCAATGTAAAGGAGATACTAGA |
| CUST_4171_PI426222305 | JHI_St_60k_v1 | DMT400061679 | CAAATTGGTATGTCTAGACAATGGTGATCAAGACGGGTTACTTATAAGTACAAAATTCCT |
Microarray Signals from CUST_4203_PI426222305
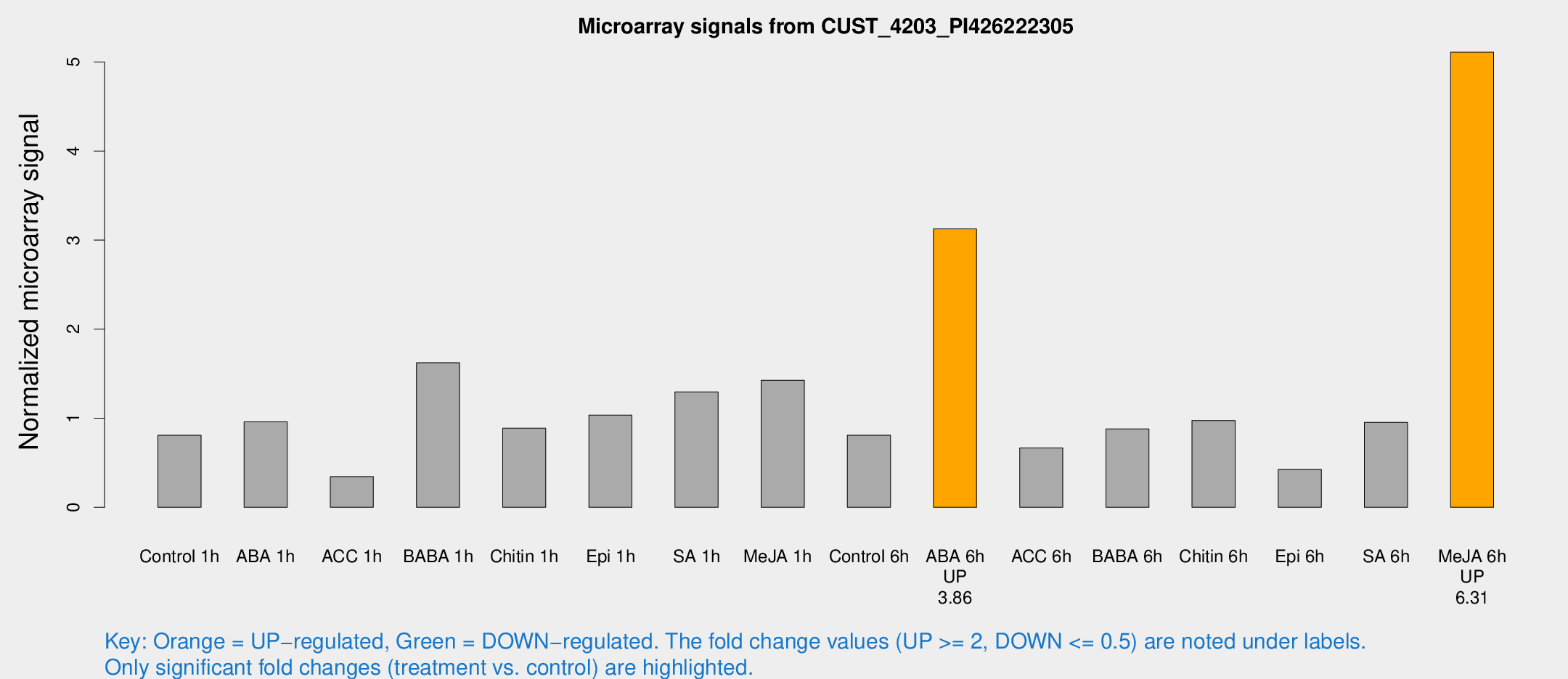
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 961.658 | 55.744 | 0.808558 | 0.0792427 |
| ABA 1h | 1331.29 | 534.054 | 0.959641 | 0.650042 |
| ACC 1h | 745.429 | 341.278 | 0.344972 | 0.587208 |
| BABA 1h | 2073.72 | 584.738 | 1.62344 | 0.428399 |
| Chitin 1h | 1158.73 | 494.724 | 0.888361 | 0.434047 |
| Epi 1h | 1182.63 | 343.804 | 1.03475 | 0.378363 |
| SA 1h | 1586.21 | 101.528 | 1.29585 | 0.0804945 |
| Me-JA 1h | 1609.48 | 641.348 | 1.4265 | 0.513097 |
| Control 6h | 1031.31 | 260.631 | 0.810085 | 0.14558 |
| ABA 6h | 3948.9 | 228.296 | 3.12601 | 0.262085 |
| ACC 6h | 909.698 | 52.7831 | 0.667035 | 0.0729161 |
| BABA 6h | 1247 | 295.402 | 0.880134 | 0.202755 |
| Chitin 6h | 1288.84 | 255.31 | 0.974469 | 0.253213 |
| Epi 6h | 583.03 | 94.7544 | 0.425037 | 0.0979562 |
| SA 6h | 1202.71 | 306.564 | 0.954365 | 0.163593 |
| Me-JA 6h | 6242.97 | 1055.64 | 5.11051 | 0.565487 |
Source Transcript PGSC0003DMT400061678 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G46760.1 | +1 | 0.0 | 556 | 290/459 (63%) | D-arabinono-1,4-lactone oxidase family protein | chr2:19213028-19215166 REVERSE LENGTH=603 |
