Probe CUST_4171_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4171_PI426222305 | JHI_St_60k_v1 | DMT400061679 | CAAATTGGTATGTCTAGACAATGGTGATCAAGACGGGTTACTTATAAGTACAAAATTCCT |
All Microarray Probes Designed to Gene DMG400024003
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4203_PI426222305 | JHI_St_60k_v1 | DMT400061678 | GCACTATCGATATCCAAACAAGGAAAACAATAAGCACATCAATGTAAAGGAGATACTAGA |
| CUST_4171_PI426222305 | JHI_St_60k_v1 | DMT400061679 | CAAATTGGTATGTCTAGACAATGGTGATCAAGACGGGTTACTTATAAGTACAAAATTCCT |
Microarray Signals from CUST_4171_PI426222305
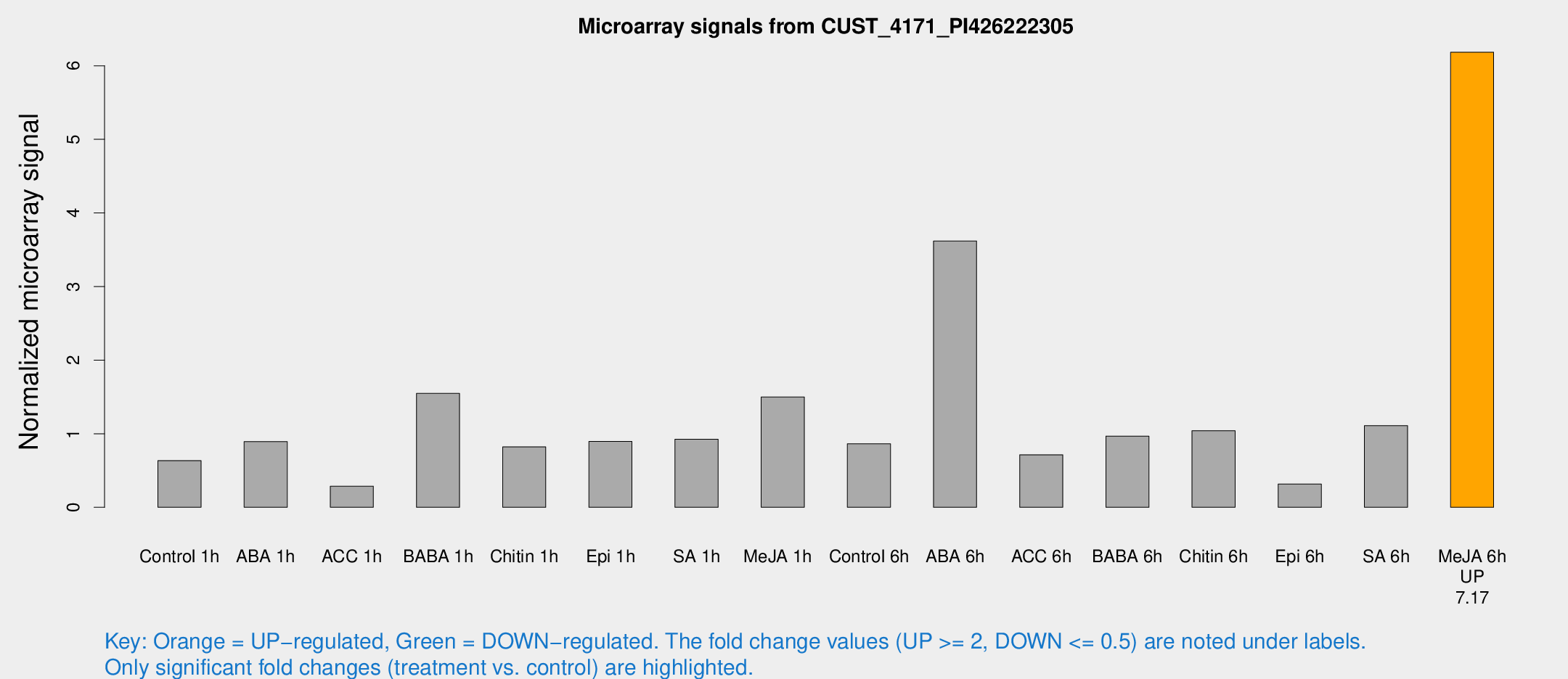
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 83.8377 | 6.44547 | 0.634969 | 0.0536281 |
| ABA 1h | 147.798 | 61.7586 | 0.892204 | 0.811367 |
| ACC 1h | 65.3344 | 29.548 | 0.286309 | 0.420519 |
| BABA 1h | 214.562 | 56.4589 | 1.5472 | 0.348034 |
| Chitin 1h | 113.496 | 44.1623 | 0.82026 | 0.332948 |
| Epi 1h | 111.053 | 29.3611 | 0.894215 | 0.291594 |
| SA 1h | 126.722 | 16.4157 | 0.924876 | 0.0640773 |
| Me-JA 1h | 212.518 | 112.705 | 1.49859 | 0.827782 |
| Control 6h | 138.505 | 53.2899 | 0.862875 | 0.372342 |
| ABA 6h | 517.802 | 76.7844 | 3.61835 | 0.431728 |
| ACC 6h | 108.446 | 10.1191 | 0.713513 | 0.0950163 |
| BABA 6h | 178.718 | 72.5671 | 0.967187 | 0.555956 |
| Chitin 6h | 148.384 | 20.2417 | 1.04157 | 0.18305 |
| Epi 6h | 48.7463 | 8.99088 | 0.316634 | 0.0928183 |
| SA 6h | 154.177 | 36.8417 | 1.10897 | 0.397378 |
| Me-JA 6h | 880.286 | 228.989 | 6.18369 | 1.44658 |
Source Transcript PGSC0003DMT400061679 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G46760.1 | +1 | 0.0 | 676 | 352/576 (61%) | D-arabinono-1,4-lactone oxidase family protein | chr2:19213028-19215166 REVERSE LENGTH=603 |
