Probe CUST_41401_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41401_PI426222305 | JHI_St_60k_v1 | DMT400079474 | GGGGTTTGTAATCTCATTTTCAGTTTCTCTTTCAGGACTTAATTATGCAGTTGTAATTGG |
All Microarray Probes Designed to Gene DMG400030946
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41401_PI426222305 | JHI_St_60k_v1 | DMT400079474 | GGGGTTTGTAATCTCATTTTCAGTTTCTCTTTCAGGACTTAATTATGCAGTTGTAATTGG |
Microarray Signals from CUST_41401_PI426222305
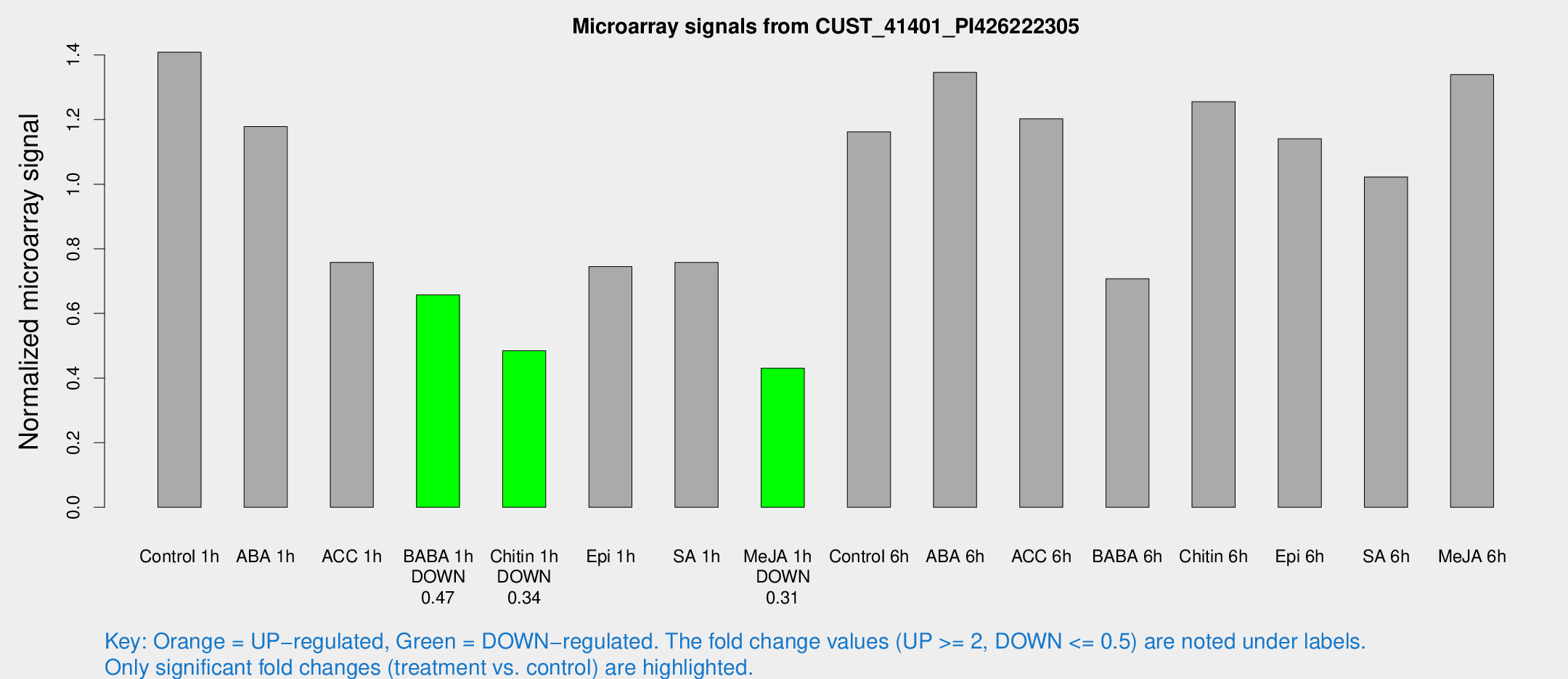
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5089.02 | 294.035 | 1.4085 | 0.0813253 |
| ABA 1h | 3781.08 | 262.296 | 1.17824 | 0.098184 |
| ACC 1h | 3260.5 | 1037.89 | 0.758162 | 0.284049 |
| BABA 1h | 2380.5 | 473.418 | 0.657696 | 0.0821833 |
| Chitin 1h | 1630.65 | 319.113 | 0.48443 | 0.0699755 |
| Epi 1h | 2407.81 | 440.109 | 0.744804 | 0.139218 |
| SA 1h | 3010.12 | 799.358 | 0.757951 | 0.191109 |
| Me-JA 1h | 1278.31 | 104.694 | 0.430564 | 0.0248892 |
| Control 6h | 4366.57 | 885.672 | 1.16204 | 0.209452 |
| ABA 6h | 5176.27 | 299.413 | 1.34597 | 0.0777156 |
| ACC 6h | 5017 | 437.602 | 1.20236 | 0.210845 |
| BABA 6h | 2958.26 | 559.362 | 0.707585 | 0.136726 |
| Chitin 6h | 4899.88 | 596.823 | 1.25509 | 0.202165 |
| Epi 6h | 4684.67 | 423.675 | 1.14061 | 0.147887 |
| SA 6h | 3824.88 | 753.93 | 1.02227 | 0.110277 |
| Me-JA 6h | 5239.2 | 1524.08 | 1.33927 | 0.366779 |
Source Transcript PGSC0003DMT400079474 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G34590.1 | +1 | 4e-28 | 109 | 75/139 (54%) | G-box binding factor 6 | chr4:16522449-16522928 FORWARD LENGTH=159 |
