Probe CUST_41381_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41381_PI426222305 | JHI_St_60k_v1 | DMT400079507 | CTCTTCTTCTTCATATGATGAAGATTCAATGTGATGTTTGTGATAAAGAAGAGGCATCAG |
All Microarray Probes Designed to Gene DMG400030958
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41381_PI426222305 | JHI_St_60k_v1 | DMT400079507 | CTCTTCTTCTTCATATGATGAAGATTCAATGTGATGTTTGTGATAAAGAAGAGGCATCAG |
| CUST_41424_PI426222305 | JHI_St_60k_v1 | DMT400079508 | GTCAAATCACTTGGACTTGTTGCAATATGTTAATCTGAATGGGAATGGGTTTTAACAGAT |
Microarray Signals from CUST_41381_PI426222305
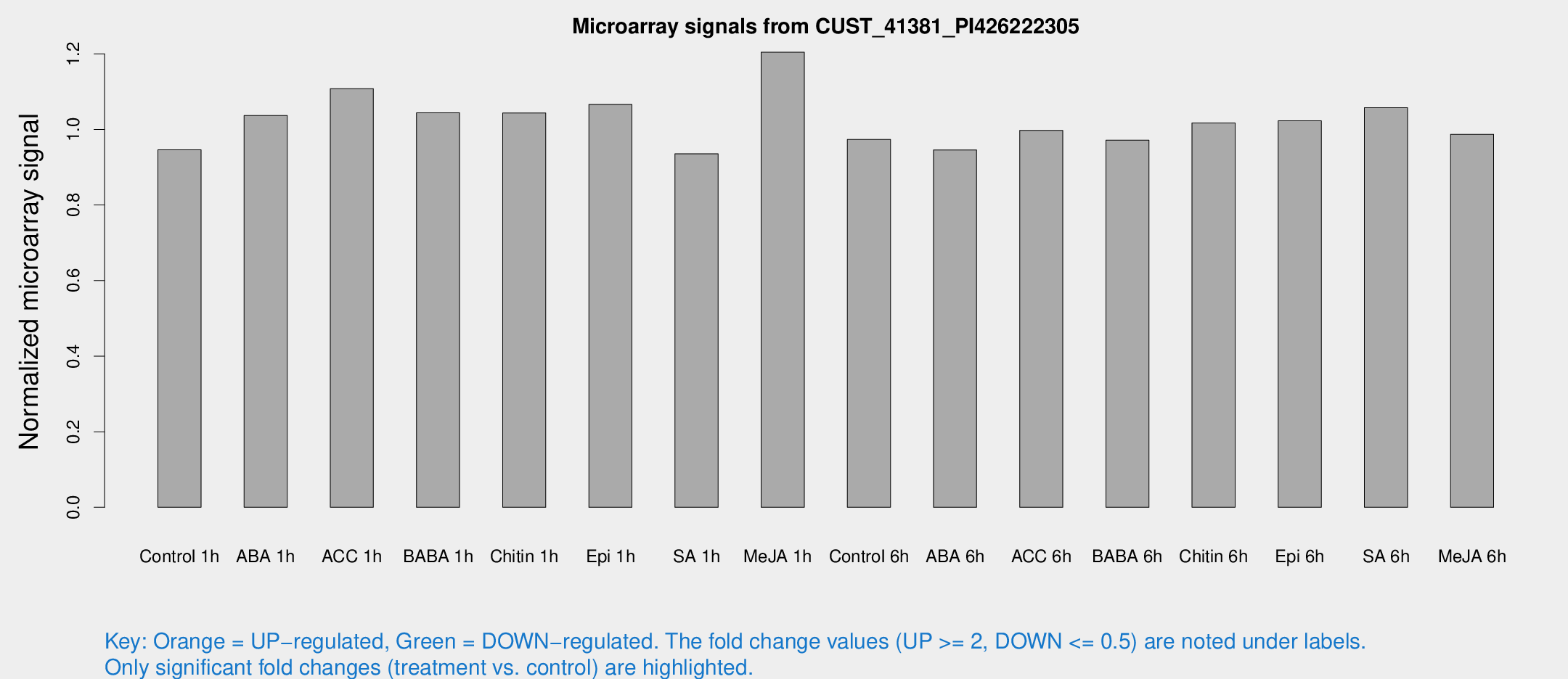
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.86566 | 3.40099 | 0.945947 | 0.547753 |
| ABA 1h | 5.68895 | 3.30151 | 1.03694 | 0.60089 |
| ACC 1h | 7.25394 | 4.33126 | 1.10789 | 0.641555 |
| BABA 1h | 6.23908 | 3.6189 | 1.04404 | 0.604924 |
| Chitin 1h | 5.8299 | 3.38744 | 1.04333 | 0.604178 |
| Epi 1h | 5.72656 | 3.32405 | 1.06618 | 0.617669 |
| SA 1h | 5.95392 | 3.44945 | 0.935461 | 0.541711 |
| Me-JA 1h | 6.10439 | 3.54392 | 1.20431 | 0.697313 |
| Control 6h | 6.04458 | 3.5013 | 0.973599 | 0.563807 |
| ABA 6h | 6.2327 | 3.61301 | 0.945864 | 0.54784 |
| ACC 6h | 7.26173 | 4.31569 | 0.997299 | 0.57754 |
| BABA 6h | 6.7481 | 3.92064 | 0.971311 | 0.563181 |
| Chitin 6h | 6.70609 | 3.88375 | 1.01715 | 0.589005 |
| Epi 6h | 7.21215 | 4.21799 | 1.02295 | 0.592345 |
| SA 6h | 6.48399 | 3.75764 | 1.05766 | 0.612473 |
| Me-JA 6h | 6.09057 | 3.52878 | 0.987053 | 0.571563 |
Source Transcript PGSC0003DMT400079507 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G39070.1 | +1 | 4e-76 | 234 | 123/208 (59%) | B-box zinc finger family protein | chr4:18205061-18206421 REVERSE LENGTH=242 |
