Probe CUST_4116_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4116_PI426222305 | JHI_St_60k_v1 | DMT400008432 | CGAAACACTAAAGTTTCTCAAGGCAAACAAGGAAGGAAAGCCAGAGAAATTCGTTAAAGA |
All Microarray Probes Designed to Gene DMG400003257
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4073_PI426222305 | JHI_St_60k_v1 | DMT400008433 | AGTGTATGTTATGAATCTTGACAGTTTCTGACCACACGTACATAAGAAAGCTCTCACCAT |
| CUST_4116_PI426222305 | JHI_St_60k_v1 | DMT400008432 | CGAAACACTAAAGTTTCTCAAGGCAAACAAGGAAGGAAAGCCAGAGAAATTCGTTAAAGA |
Microarray Signals from CUST_4116_PI426222305
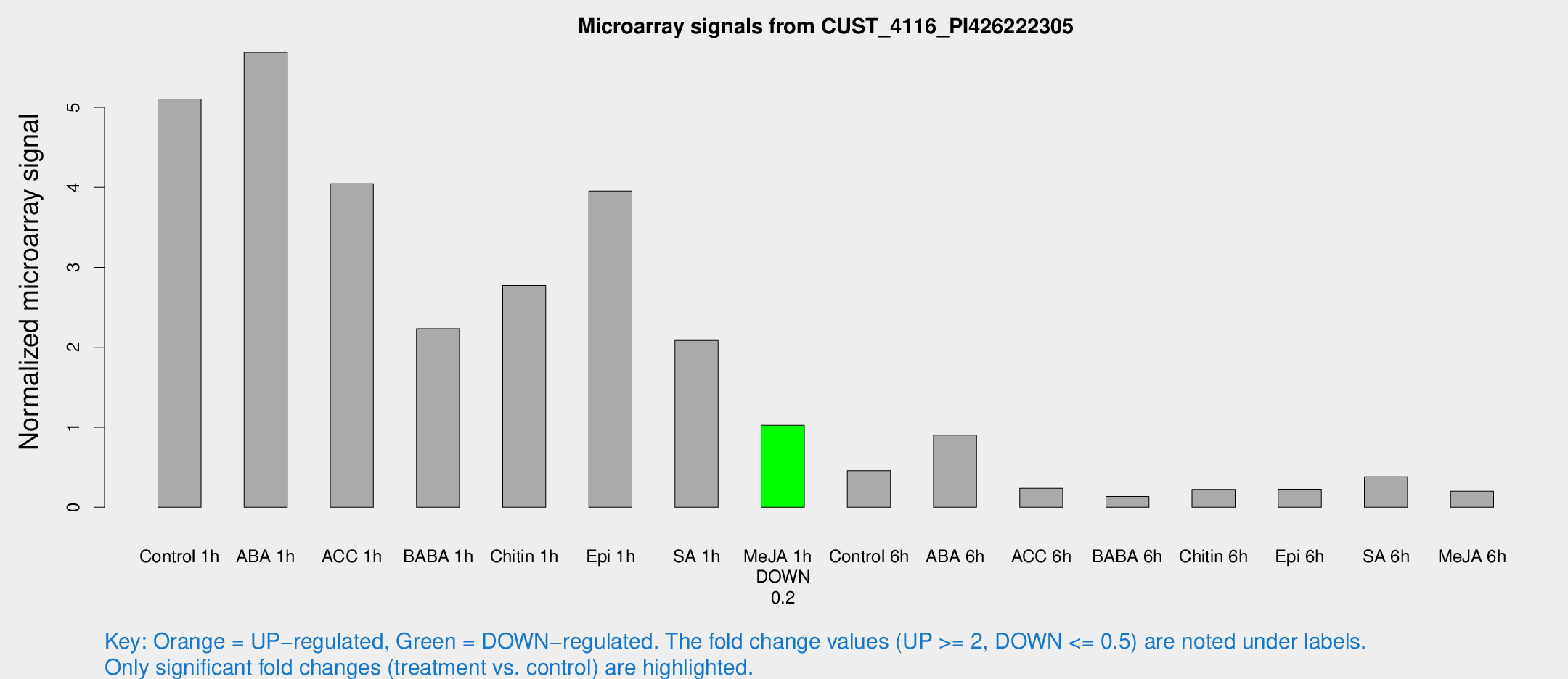
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 221.552 | 18.4538 | 5.10361 | 0.304059 |
| ABA 1h | 239.008 | 69.2565 | 5.68856 | 2.36156 |
| ACC 1h | 181.707 | 21.3118 | 4.0443 | 0.694332 |
| BABA 1h | 111.222 | 37.5778 | 2.23307 | 0.90437 |
| Chitin 1h | 107.894 | 7.00522 | 2.77291 | 0.179408 |
| Epi 1h | 149.585 | 17.5104 | 3.95458 | 0.562894 |
| SA 1h | 101.763 | 31 | 2.08617 | 0.550296 |
| Me-JA 1h | 36.7123 | 4.56554 | 1.02535 | 0.223511 |
| Control 6h | 23.723 | 8.94356 | 0.45876 | 0.187263 |
| ABA 6h | 46.6781 | 14.8253 | 0.902776 | 0.327021 |
| ACC 6h | 17.476 | 11.1455 | 0.235397 | 0.13775 |
| BABA 6h | 6.48397 | 3.67916 | 0.133702 | 0.075924 |
| Chitin 6h | 12.9738 | 6.55049 | 0.223262 | 0.114033 |
| Epi 6h | 12.4318 | 4.17463 | 0.225544 | 0.0911637 |
| SA 6h | 17.2662 | 4.42693 | 0.380978 | 0.0935719 |
| Me-JA 6h | 9.09777 | 3.367 | 0.199155 | 0.0829545 |
Source Transcript PGSC0003DMT400008432 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G36110.1 | +2 | 8e-154 | 454 | 221/479 (46%) | cytochrome P450, family 716, subfamily A, polypeptide 1 | chr5:14195377-14197613 FORWARD LENGTH=477 |
