Probe CUST_4110_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4110_PI426222305 | JHI_St_60k_v1 | DMT400038256 | AACCATCAAGATTTGAAGAAGCTGGACCTGCACCATATATAGATGCTCGTCTTTCTGCAC |
All Microarray Probes Designed to Gene DMG400014760
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4110_PI426222305 | JHI_St_60k_v1 | DMT400038256 | AACCATCAAGATTTGAAGAAGCTGGACCTGCACCATATATAGATGCTCGTCTTTCTGCAC |
Microarray Signals from CUST_4110_PI426222305
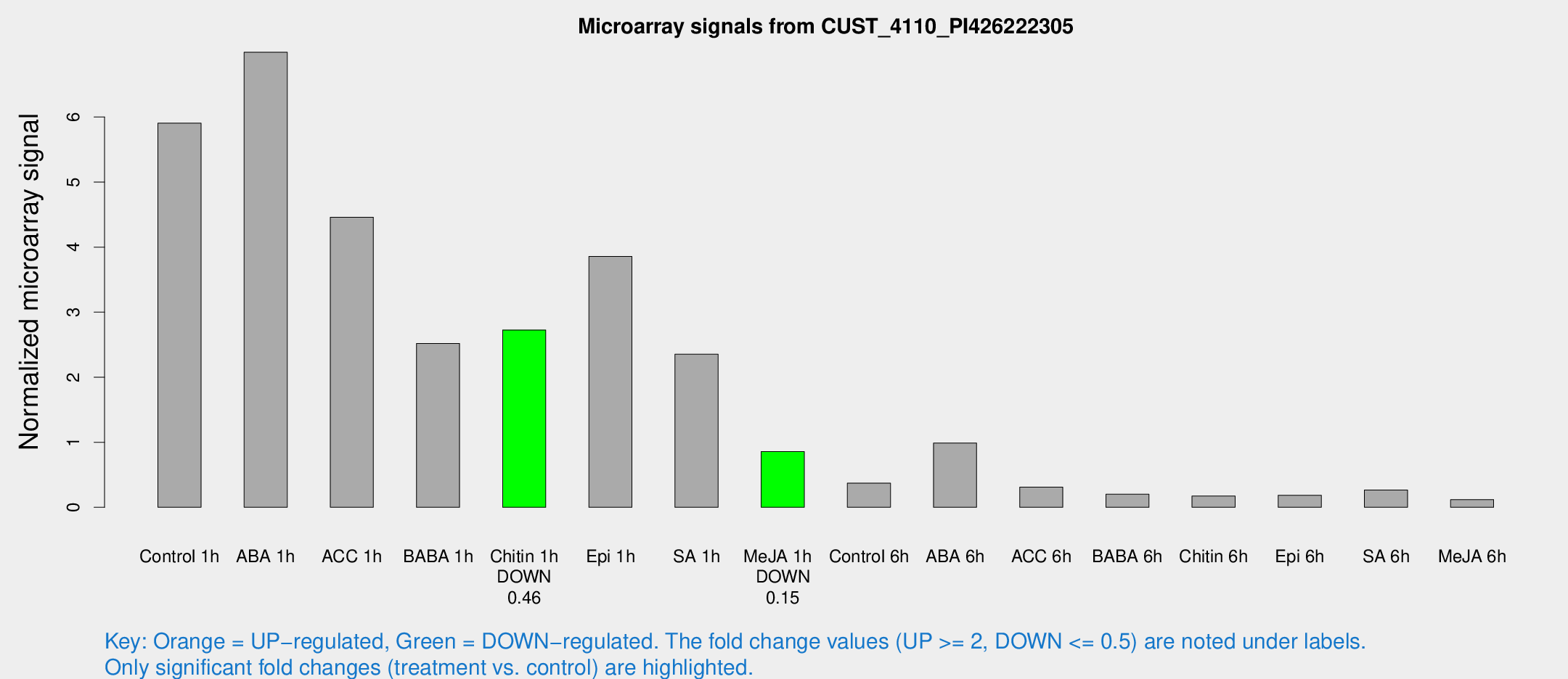
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 933.542 | 179.689 | 5.90697 | 0.875599 |
| ABA 1h | 1013.96 | 279.668 | 6.99653 | 2.43174 |
| ACC 1h | 724.607 | 153.092 | 4.45937 | 0.986769 |
| BABA 1h | 448.965 | 156.817 | 2.51944 | 1.10676 |
| Chitin 1h | 390.596 | 90.4901 | 2.72585 | 0.399945 |
| Epi 1h | 507.706 | 29.5579 | 3.85711 | 0.223948 |
| SA 1h | 416.629 | 156.688 | 2.35393 | 0.731633 |
| Me-JA 1h | 111.279 | 21.8659 | 0.857942 | 0.136276 |
| Control 6h | 60.0967 | 15.8447 | 0.371325 | 0.0907166 |
| ABA 6h | 190.264 | 63.8797 | 0.988182 | 0.479658 |
| ACC 6h | 53.99 | 4.94557 | 0.30966 | 0.0567805 |
| BABA 6h | 34.6259 | 4.04023 | 0.201227 | 0.0320332 |
| Chitin 6h | 33.1095 | 14.1719 | 0.173253 | 0.0707649 |
| Epi 6h | 37.6619 | 12.9935 | 0.184635 | 0.0801696 |
| SA 6h | 42.0932 | 10.1795 | 0.264805 | 0.0415739 |
| Me-JA 6h | 18.7981 | 4.28974 | 0.118547 | 0.0232627 |
Source Transcript PGSC0003DMT400038256 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G36110.1 | +1 | 1e-45 | 159 | 79/157 (50%) | cytochrome P450, family 716, subfamily A, polypeptide 1 | chr5:14195377-14197613 FORWARD LENGTH=477 |
