Probe CUST_40535_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40535_PI426222305 | JHI_St_60k_v1 | DMT400029949 | TGTGGTATCTTGAAAACTACTCGATACAACCTAACGATACATCCAATACCAATGATTGTG |
All Microarray Probes Designed to Gene DMG400011499
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40495_PI426222305 | JHI_St_60k_v1 | DMT400029947 | GGAACTTAACAAAATCAAAGGGCTTAAAGTTGATGTTGCTACTGTAGAGACTAACATTGT |
| CUST_40535_PI426222305 | JHI_St_60k_v1 | DMT400029949 | TGTGGTATCTTGAAAACTACTCGATACAACCTAACGATACATCCAATACCAATGATTGTG |
Microarray Signals from CUST_40535_PI426222305
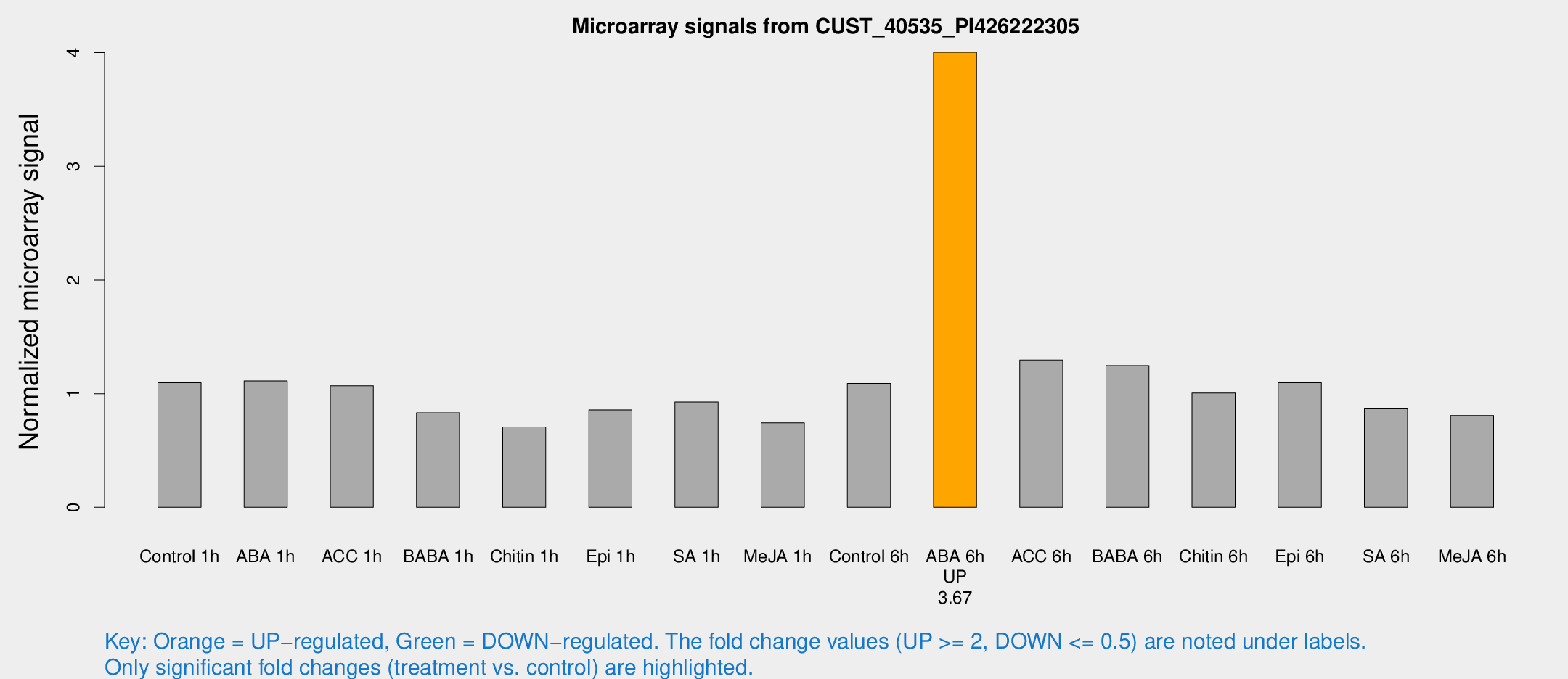
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 336.859 | 41.7917 | 1.09646 | 0.0668126 |
| ABA 1h | 298.761 | 17.5981 | 1.11221 | 0.101665 |
| ACC 1h | 350.529 | 72.6038 | 1.06927 | 0.176625 |
| BABA 1h | 245.312 | 24.6391 | 0.831416 | 0.0495592 |
| Chitin 1h | 196.127 | 27.0868 | 0.707368 | 0.0426779 |
| Epi 1h | 228.205 | 28.3186 | 0.857601 | 0.0846216 |
| SA 1h | 289.346 | 17.0744 | 0.927358 | 0.0643323 |
| Me-JA 1h | 184.408 | 11.2139 | 0.744321 | 0.0545463 |
| Control 6h | 358.922 | 89.5819 | 1.08975 | 0.234932 |
| ABA 6h | 1296.98 | 100.199 | 4.00381 | 0.240847 |
| ACC 6h | 459.065 | 63.4222 | 1.29567 | 0.0832883 |
| BABA 6h | 430.088 | 58.0998 | 1.24725 | 0.122138 |
| Chitin 6h | 325.699 | 20.7104 | 1.00651 | 0.0699496 |
| Epi 6h | 375.073 | 22.0739 | 1.0972 | 0.0934594 |
| SA 6h | 267.584 | 48.8835 | 0.866371 | 0.133577 |
| Me-JA 6h | 247.388 | 30.6094 | 0.80836 | 0.0827391 |
Source Transcript PGSC0003DMT400029949 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G08630.1 | +1 | 5e-180 | 518 | 250/354 (71%) | threonine aldolase 1 | chr1:2743948-2745685 REVERSE LENGTH=358 |
