Probe CUST_40495_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40495_PI426222305 | JHI_St_60k_v1 | DMT400029947 | GGAACTTAACAAAATCAAAGGGCTTAAAGTTGATGTTGCTACTGTAGAGACTAACATTGT |
All Microarray Probes Designed to Gene DMG400011499
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40495_PI426222305 | JHI_St_60k_v1 | DMT400029947 | GGAACTTAACAAAATCAAAGGGCTTAAAGTTGATGTTGCTACTGTAGAGACTAACATTGT |
| CUST_40535_PI426222305 | JHI_St_60k_v1 | DMT400029949 | TGTGGTATCTTGAAAACTACTCGATACAACCTAACGATACATCCAATACCAATGATTGTG |
Microarray Signals from CUST_40495_PI426222305
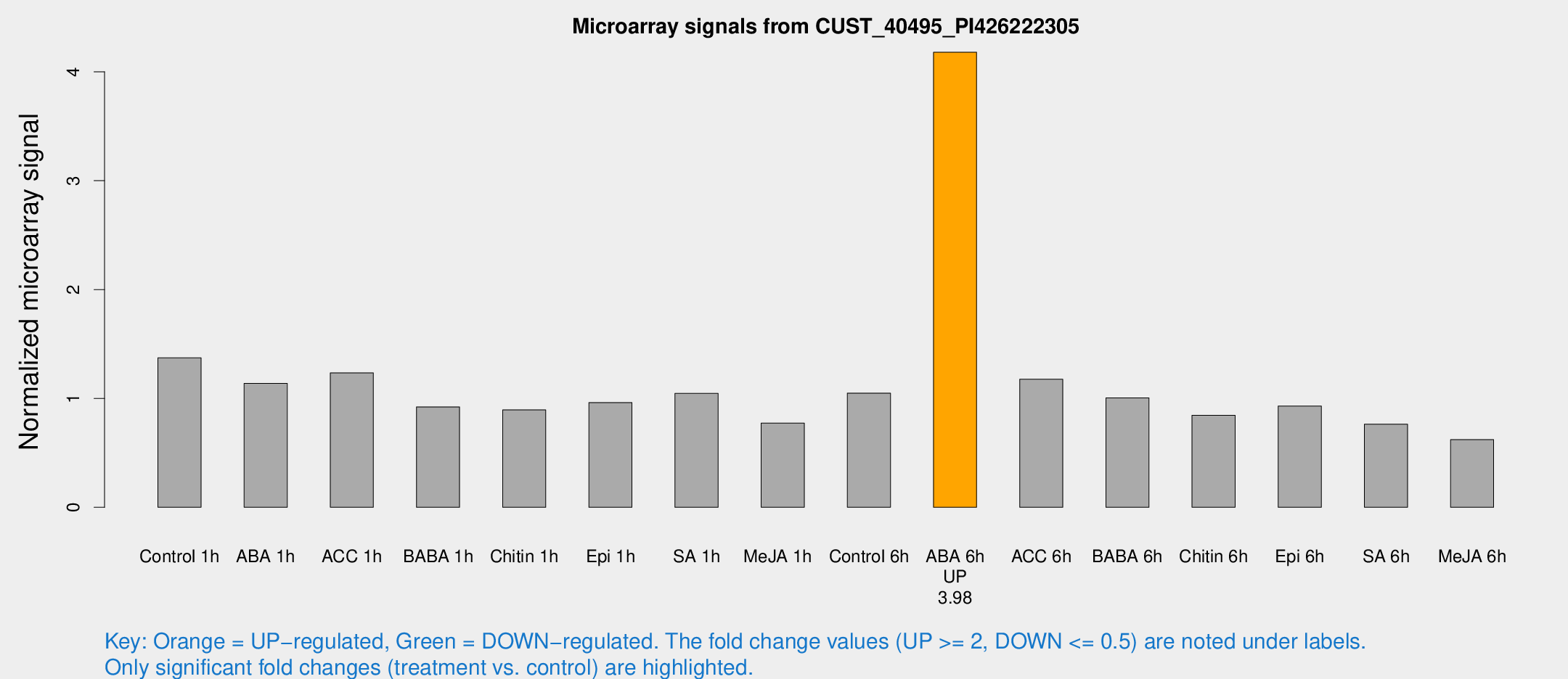
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 934.921 | 163.696 | 1.37236 | 0.143796 |
| ABA 1h | 666.677 | 38.6048 | 1.13871 | 0.090282 |
| ACC 1h | 901.165 | 218.26 | 1.23442 | 0.255539 |
| BABA 1h | 606.543 | 98.6877 | 0.922151 | 0.0690501 |
| Chitin 1h | 540.803 | 65.1802 | 0.894954 | 0.051923 |
| Epi 1h | 568.138 | 98.8852 | 0.96228 | 0.146358 |
| SA 1h | 722.091 | 88.8756 | 1.04623 | 0.112617 |
| Me-JA 1h | 420.377 | 30.4851 | 0.773886 | 0.0532927 |
| Control 6h | 797.265 | 248.038 | 1.04893 | 0.347579 |
| ABA 6h | 2978.73 | 323.104 | 4.17965 | 0.31472 |
| ACC 6h | 901.335 | 90.7834 | 1.17579 | 0.068057 |
| BABA 6h | 750.497 | 63.4371 | 1.00551 | 0.0582511 |
| Chitin 6h | 599.486 | 53.2227 | 0.844636 | 0.0788536 |
| Epi 6h | 709.08 | 103.286 | 0.929504 | 0.191056 |
| SA 6h | 530.688 | 125.138 | 0.763462 | 0.117341 |
| Me-JA 6h | 418.184 | 60.4892 | 0.621281 | 0.0383428 |
Source Transcript PGSC0003DMT400029947 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G08630.1 | +1 | 8e-163 | 461 | 223/291 (77%) | threonine aldolase 1 | chr1:2743948-2745685 REVERSE LENGTH=358 |
