Probe CUST_40327_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40327_PI426222305 | JHI_St_60k_v1 | DMT400048375 | AGAAGCTAATCAAGGCCCATTCTTTGGTTTTATTGCCAAGAATGAAATCTCTAATGGAAG |
All Microarray Probes Designed to Gene DMG400018793
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40327_PI426222305 | JHI_St_60k_v1 | DMT400048375 | AGAAGCTAATCAAGGCCCATTCTTTGGTTTTATTGCCAAGAATGAAATCTCTAATGGAAG |
| CUST_40335_PI426222305 | JHI_St_60k_v1 | DMT400048376 | AGAAGCTAATCAAGGCCCATTCTTTGGTTTTATTGCCAAGAATGAAATCTCTAATGGAAG |
Microarray Signals from CUST_40327_PI426222305
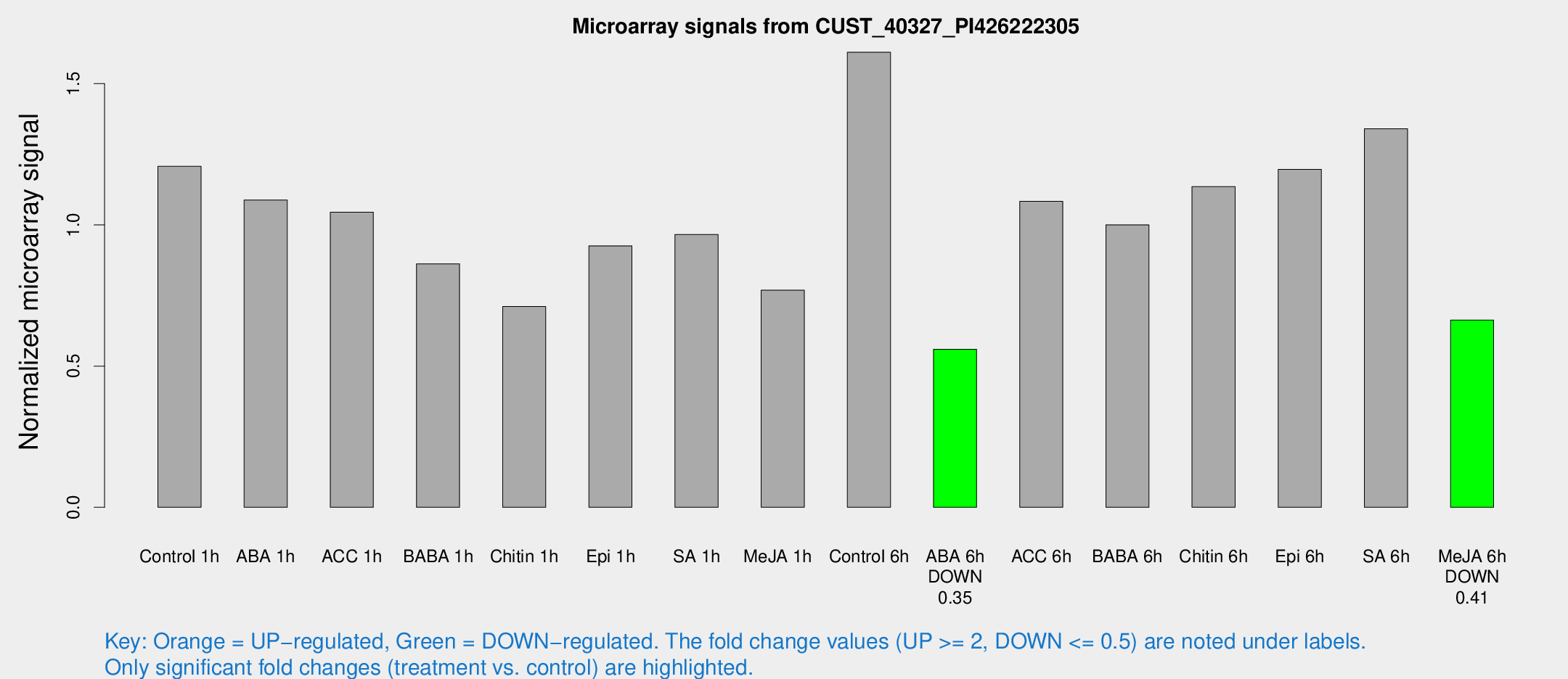
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2496.68 | 443.661 | 1.20741 | 0.130406 |
| ABA 1h | 1939.62 | 133.541 | 1.08809 | 0.120367 |
| ACC 1h | 2171.67 | 208.367 | 1.04491 | 0.0814617 |
| BABA 1h | 1730.26 | 309.298 | 0.862059 | 0.0823829 |
| Chitin 1h | 1289.7 | 93.5005 | 0.71107 | 0.041101 |
| Epi 1h | 1609.99 | 93.1754 | 0.925743 | 0.053487 |
| SA 1h | 1999.22 | 128.096 | 0.966063 | 0.0558016 |
| Me-JA 1h | 1270.6 | 123.183 | 0.768652 | 0.0444325 |
| Control 6h | 3381.76 | 683.326 | 1.61134 | 0.230993 |
| ABA 6h | 1198.91 | 82.4048 | 0.559453 | 0.0657108 |
| ACC 6h | 2576.42 | 476.118 | 1.08331 | 0.0625734 |
| BABA 6h | 2316.35 | 406.986 | 1.00007 | 0.197159 |
| Chitin 6h | 2485.21 | 405.51 | 1.13551 | 0.143748 |
| Epi 6h | 2707.56 | 156.423 | 1.19677 | 0.112871 |
| SA 6h | 2773.63 | 525.952 | 1.34053 | 0.128153 |
| Me-JA 6h | 1337.47 | 136.63 | 0.662976 | 0.0383216 |
Source Transcript PGSC0003DMT400048375 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G34000.1 | +2 | 8e-35 | 89 | 63/122 (52%) | one-helix protein 2 | chr1:12358151-12358902 REVERSE LENGTH=172 |
