Probe CUST_39155_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39155_PI426222305 | JHI_St_60k_v1 | DMT400036325 | GTAGAGCACGGAAGAGGTTCGGCGATGGTTACTGGTAACACTGCTTCACTTTTCTTTTTT |
All Microarray Probes Designed to Gene DMG400013973
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39140_PI426222305 | JHI_St_60k_v1 | DMT400036326 | GCTTTCTCAGGAATGCTTTTTTAGTGTTGAGCTGTGTTTGTCTTGTCCTTGTAAATAATT |
| CUST_39155_PI426222305 | JHI_St_60k_v1 | DMT400036325 | GTAGAGCACGGAAGAGGTTCGGCGATGGTTACTGGTAACACTGCTTCACTTTTCTTTTTT |
Microarray Signals from CUST_39155_PI426222305
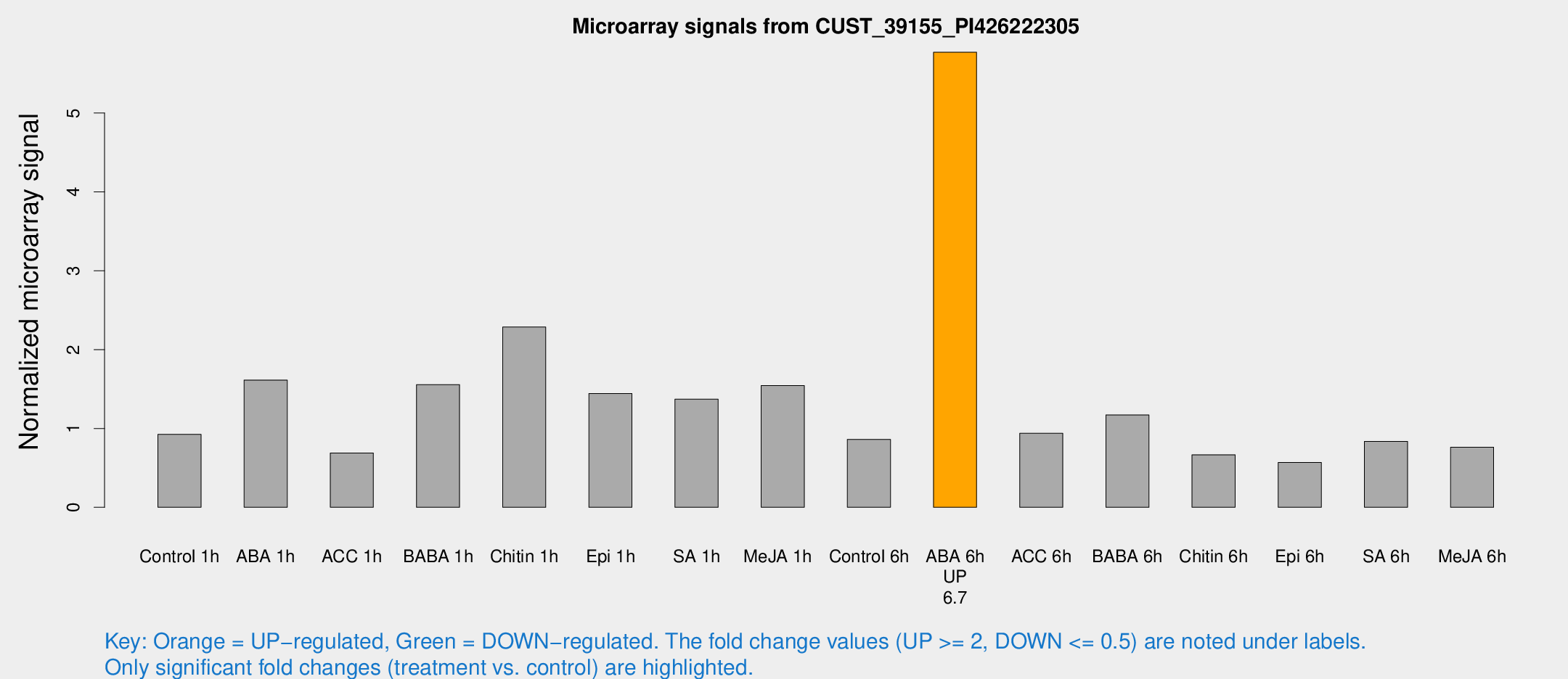
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 114.508 | 13.6086 | 0.925996 | 0.159498 |
| ABA 1h | 202.478 | 79.242 | 1.61402 | 0.548422 |
| ACC 1h | 89.3359 | 17.2481 | 0.689042 | 0.115389 |
| BABA 1h | 201.466 | 57.3175 | 1.55595 | 0.362527 |
| Chitin 1h | 254.256 | 29.906 | 2.28603 | 0.407561 |
| Epi 1h | 163.398 | 38.9306 | 1.44371 | 0.388808 |
| SA 1h | 185.38 | 49.4967 | 1.3723 | 0.368094 |
| Me-JA 1h | 157.392 | 24.2307 | 1.54463 | 0.173718 |
| Control 6h | 111.859 | 28.6027 | 0.862023 | 0.20471 |
| ABA 6h | 771.103 | 127.971 | 5.77131 | 0.708323 |
| ACC 6h | 132.393 | 12.0044 | 0.938539 | 0.0624138 |
| BABA 6h | 170.94 | 45.6316 | 1.17139 | 0.263237 |
| Chitin 6h | 88.2778 | 12.8146 | 0.666514 | 0.0954631 |
| Epi 6h | 88.7271 | 29.5964 | 0.569768 | 0.172907 |
| SA 6h | 103.805 | 16.2368 | 0.836266 | 0.0566716 |
| Me-JA 6h | 101.261 | 27.1108 | 0.762397 | 0.200702 |
Source Transcript PGSC0003DMT400036325 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
