Probe CUST_39145_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39145_PI426222305 | JHI_St_60k_v1 | DMT400035577 | GGGGAAGAATGGTTAGATTCTACTTCTTTTCCTTATGATAATGGGGTGTAATTAACTAGT |
All Microarray Probes Designed to Gene DMG400013680
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39145_PI426222305 | JHI_St_60k_v1 | DMT400035577 | GGGGAAGAATGGTTAGATTCTACTTCTTTTCCTTATGATAATGGGGTGTAATTAACTAGT |
| CUST_39122_PI426222305 | JHI_St_60k_v1 | DMT400035576 | CCTTCCGCTAAATCTGAACAAGTCGAAAGAAATAAACCCATAACGTGATTTTTACAATTG |
Microarray Signals from CUST_39145_PI426222305
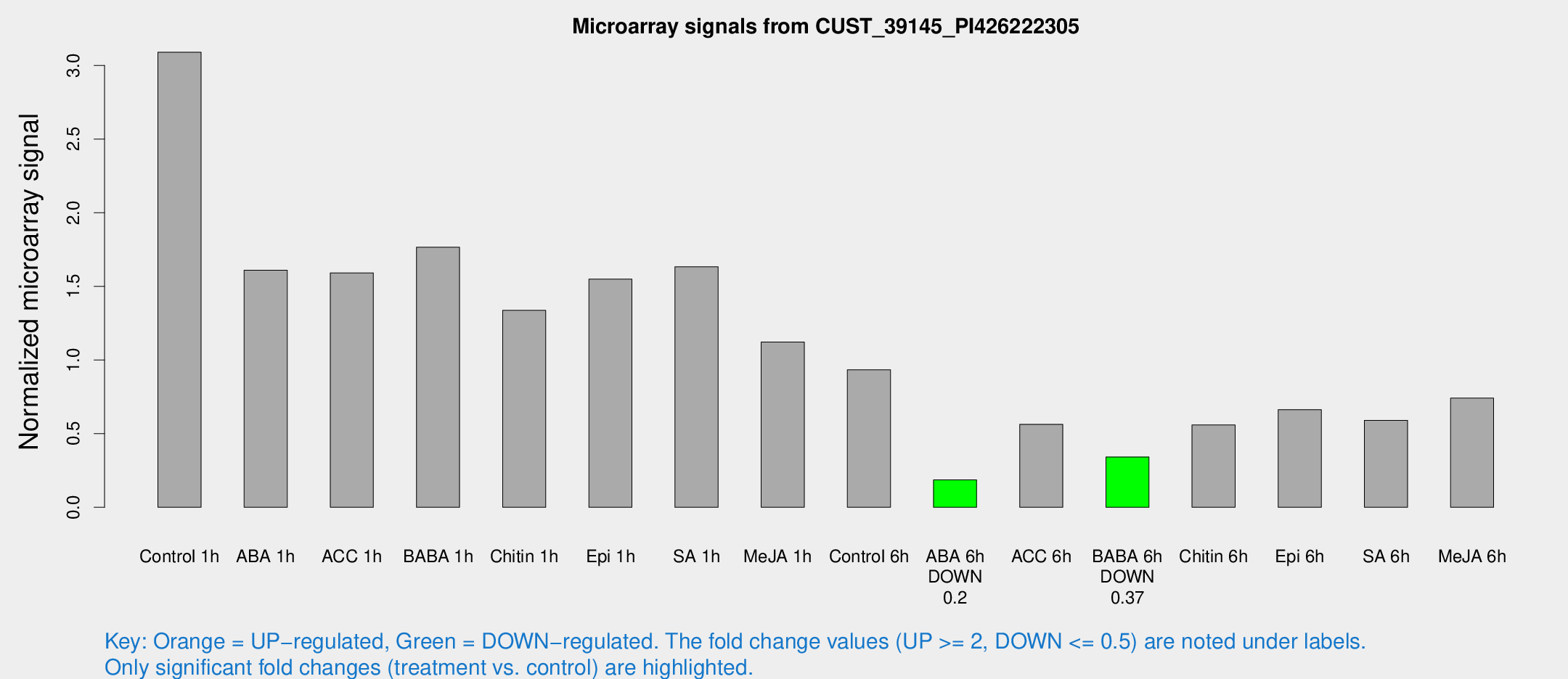
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11139.9 | 2057.68 | 3.0895 | 0.360658 |
| ABA 1h | 5076.02 | 702.499 | 1.60971 | 0.120933 |
| ACC 1h | 6416.77 | 1887.02 | 1.59167 | 0.491613 |
| BABA 1h | 6338.06 | 1471.68 | 1.76539 | 0.290198 |
| Chitin 1h | 4278.51 | 592.288 | 1.33724 | 0.0822897 |
| Epi 1h | 4779.44 | 650.631 | 1.5487 | 0.213249 |
| SA 1h | 6145.26 | 1216.7 | 1.63244 | 0.291832 |
| Me-JA 1h | 3265.57 | 473.256 | 1.12203 | 0.065947 |
| Control 6h | 3347.31 | 515.435 | 0.933483 | 0.0975011 |
| ABA 6h | 707.174 | 106.962 | 0.186288 | 0.0171573 |
| ACC 6h | 2324.11 | 368.023 | 0.563434 | 0.0893948 |
| BABA 6h | 1386.02 | 269.23 | 0.341974 | 0.0675208 |
| Chitin 6h | 2106.26 | 235.852 | 0.559323 | 0.0529062 |
| Epi 6h | 2617.69 | 151.525 | 0.662842 | 0.0382814 |
| SA 6h | 2240.08 | 596.912 | 0.59027 | 0.125704 |
| Me-JA 6h | 2683.4 | 556.539 | 0.741184 | 0.0943026 |
Source Transcript PGSC0003DMT400035577 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G26440.1 | +2 | 0.0 | 627 | 337/536 (63%) | Plant invertase/pectin methylesterase inhibitor superfamily | chr2:11247407-11249407 FORWARD LENGTH=547 |
