Probe CUST_39122_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39122_PI426222305 | JHI_St_60k_v1 | DMT400035576 | CCTTCCGCTAAATCTGAACAAGTCGAAAGAAATAAACCCATAACGTGATTTTTACAATTG |
All Microarray Probes Designed to Gene DMG400013680
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39145_PI426222305 | JHI_St_60k_v1 | DMT400035577 | GGGGAAGAATGGTTAGATTCTACTTCTTTTCCTTATGATAATGGGGTGTAATTAACTAGT |
| CUST_39122_PI426222305 | JHI_St_60k_v1 | DMT400035576 | CCTTCCGCTAAATCTGAACAAGTCGAAAGAAATAAACCCATAACGTGATTTTTACAATTG |
Microarray Signals from CUST_39122_PI426222305
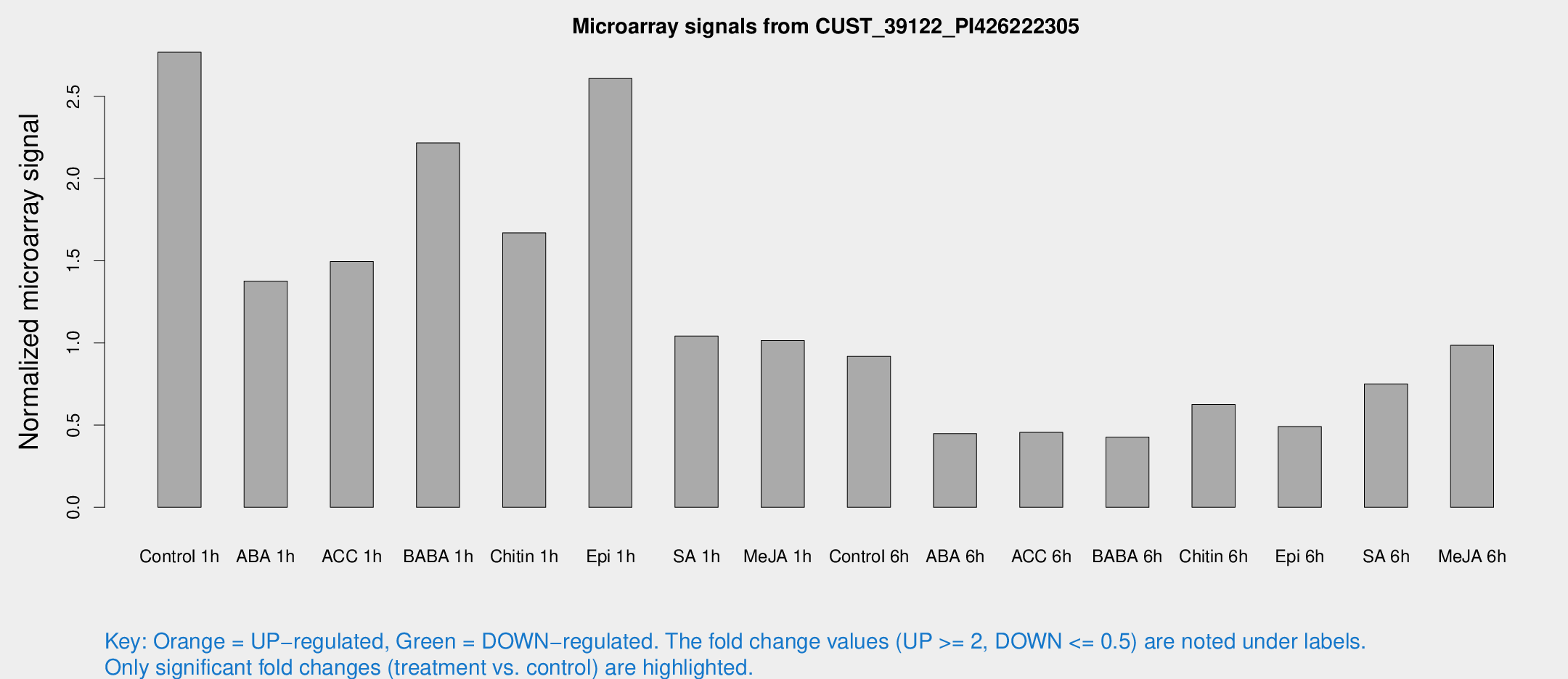
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 35.3019 | 5.4108 | 2.76855 | 0.316696 |
| ABA 1h | 15.8995 | 3.56534 | 1.37671 | 0.333364 |
| ACC 1h | 21.5022 | 6.1695 | 1.4953 | 0.460771 |
| BABA 1h | 30.575 | 9.50948 | 2.21663 | 0.705343 |
| Chitin 1h | 20.4764 | 5.62723 | 1.66978 | 0.409653 |
| Epi 1h | 28.9506 | 4.57672 | 2.60896 | 0.429294 |
| SA 1h | 16.7281 | 7.0637 | 1.04184 | 0.57999 |
| Me-JA 1h | 11.3369 | 3.64117 | 1.01416 | 0.329438 |
| Control 6h | 15.977 | 9.26781 | 0.918109 | 0.584299 |
| ABA 6h | 5.94914 | 3.2165 | 0.447857 | 0.243523 |
| ACC 6h | 6.63503 | 3.72009 | 0.456011 | 0.249639 |
| BABA 6h | 5.97425 | 3.46821 | 0.427312 | 0.247627 |
| Chitin 6h | 9.26656 | 3.48161 | 0.625882 | 0.293745 |
| Epi 6h | 6.9535 | 3.63056 | 0.490776 | 0.257331 |
| SA 6h | 11.6868 | 5.87062 | 0.750394 | 0.356208 |
| Me-JA 6h | 14.3237 | 4.65117 | 0.985854 | 0.394392 |
Source Transcript PGSC0003DMT400035576 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G26440.1 | +2 | 5e-81 | 263 | 165/307 (54%) | Plant invertase/pectin methylesterase inhibitor superfamily | chr2:11247407-11249407 FORWARD LENGTH=547 |
