Probe CUST_38846_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38846_PI426222305 | JHI_St_60k_v1 | DMT400015270 | GGATAGGAATAGTCCTTACTGGTATCTTGTATGTTGAGAAATCTGGCAAAATGTATATCA |
All Microarray Probes Designed to Gene DMG400005958
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38815_PI426222305 | JHI_St_60k_v1 | DMT400015269 | GGATAGGAATAGTCCTTACTGGTATCTTGTATGTTGAGAAATCTGGCAAAATGTATATCA |
| CUST_38846_PI426222305 | JHI_St_60k_v1 | DMT400015270 | GGATAGGAATAGTCCTTACTGGTATCTTGTATGTTGAGAAATCTGGCAAAATGTATATCA |
Microarray Signals from CUST_38846_PI426222305
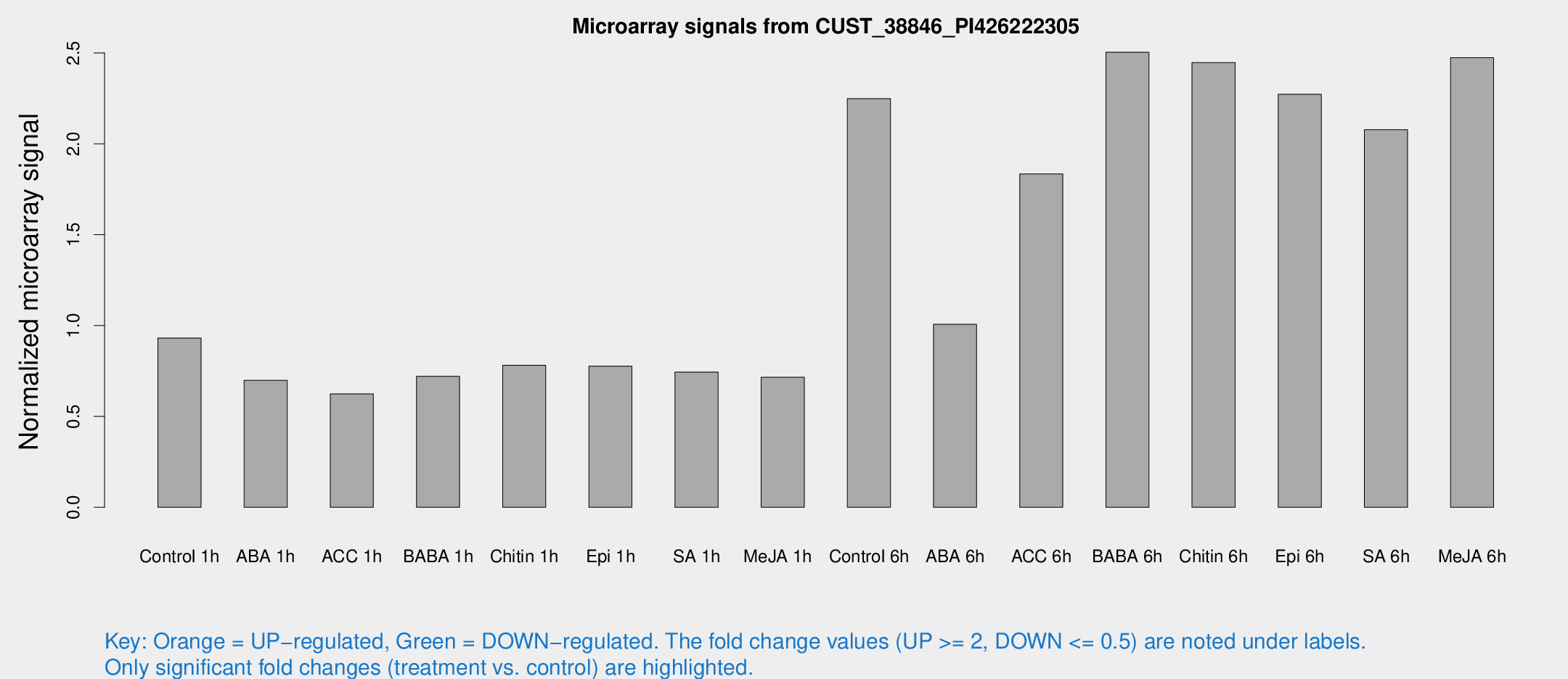
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 152.839 | 13.0943 | 0.930954 | 0.057942 |
| ABA 1h | 101.5 | 8.51541 | 0.699143 | 0.0967828 |
| ACC 1h | 114.46 | 29.956 | 0.624012 | 0.159994 |
| BABA 1h | 117.559 | 21.5593 | 0.721337 | 0.0730882 |
| Chitin 1h | 117.835 | 19.9312 | 0.781715 | 0.120924 |
| Epi 1h | 110.532 | 10.1549 | 0.776809 | 0.0574861 |
| SA 1h | 124.768 | 8.01416 | 0.743976 | 0.0477223 |
| Me-JA 1h | 95.4717 | 6.58577 | 0.715882 | 0.0637233 |
| Control 6h | 397.304 | 97.4859 | 2.24865 | 0.471404 |
| ABA 6h | 174.631 | 10.7717 | 1.00692 | 0.0871841 |
| ACC 6h | 346.922 | 36.3455 | 1.83417 | 0.154273 |
| BABA 6h | 465.774 | 62.154 | 2.50426 | 0.256834 |
| Chitin 6h | 424.889 | 24.8699 | 2.44715 | 0.143132 |
| Epi 6h | 423.063 | 49.3598 | 2.27282 | 0.261047 |
| SA 6h | 336.752 | 24.4359 | 2.07775 | 0.326488 |
| Me-JA 6h | 409.908 | 61.1237 | 2.47424 | 0.20729 |
Source Transcript PGSC0003DMT400015270 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G71010.1 | +3 | 0.0 | 1495 | 885/1746 (51%) | FORMS APLOID AND BINUCLEATE CELLS 1C | chr1:26782839-26788712 FORWARD LENGTH=1648 |
