Probe CUST_38815_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38815_PI426222305 | JHI_St_60k_v1 | DMT400015269 | GGATAGGAATAGTCCTTACTGGTATCTTGTATGTTGAGAAATCTGGCAAAATGTATATCA |
All Microarray Probes Designed to Gene DMG400005958
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38815_PI426222305 | JHI_St_60k_v1 | DMT400015269 | GGATAGGAATAGTCCTTACTGGTATCTTGTATGTTGAGAAATCTGGCAAAATGTATATCA |
| CUST_38846_PI426222305 | JHI_St_60k_v1 | DMT400015270 | GGATAGGAATAGTCCTTACTGGTATCTTGTATGTTGAGAAATCTGGCAAAATGTATATCA |
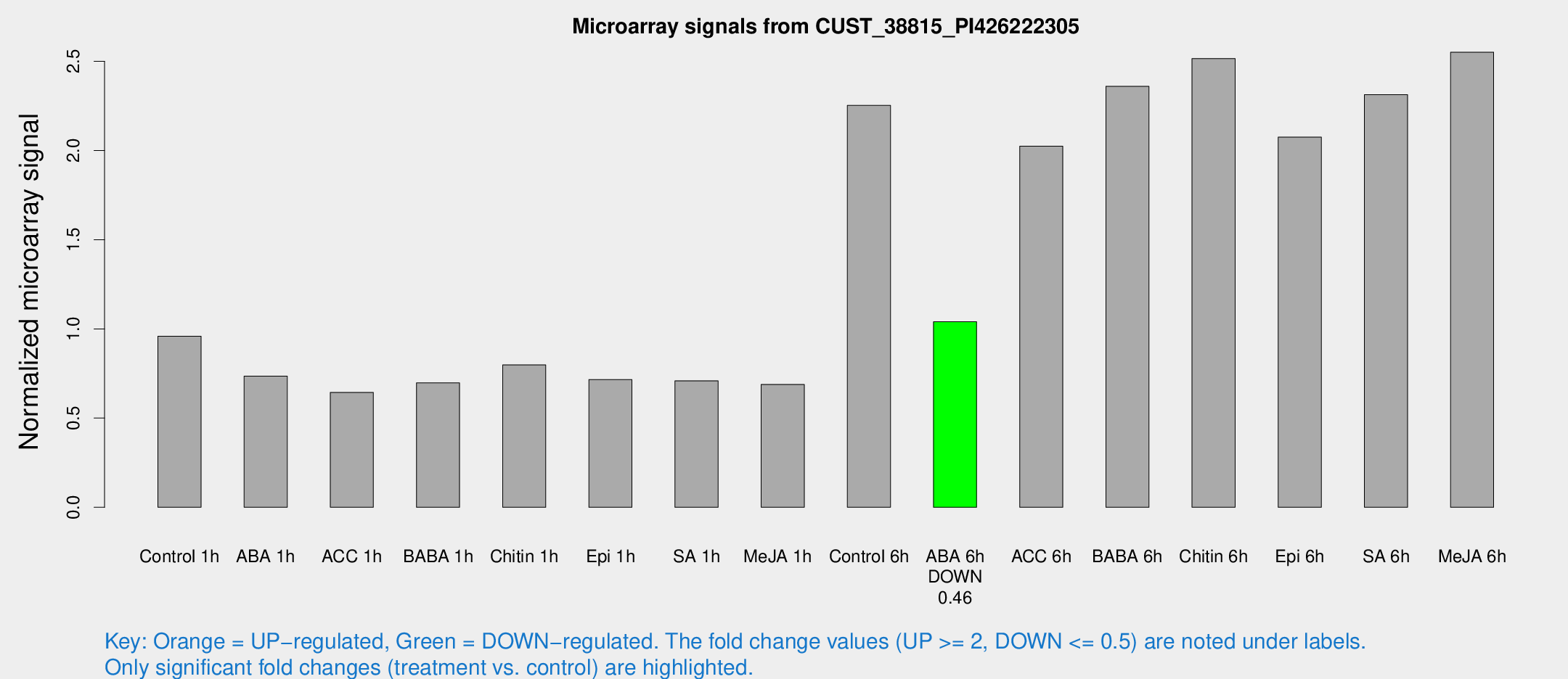
Microarray Signals from CUST_38815_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 157.736 | 9.70224 | 0.9592 | 0.058922 |
| ABA 1h | 107.516 | 8.79953 | 0.735692 | 0.048144 |
| ACC 1h | 122.742 | 38.1458 | 0.644341 | 0.206416 |
| BABA 1h | 111.726 | 12.2258 | 0.69743 | 0.0463574 |
| Chitin 1h | 119.555 | 14.1166 | 0.798236 | 0.0629618 |
| Epi 1h | 103.537 | 13.2788 | 0.716095 | 0.0874379 |
| SA 1h | 123.274 | 22.6681 | 0.709126 | 0.13569 |
| Me-JA 1h | 92.8969 | 7.70294 | 0.688705 | 0.094914 |
| Control 6h | 394.652 | 88.5643 | 2.25269 | 0.394756 |
| ABA 6h | 181.518 | 11.1065 | 1.04025 | 0.0730423 |
| ACC 6h | 383.285 | 29.3081 | 2.02426 | 0.231986 |
| BABA 6h | 439.576 | 50.6928 | 2.35985 | 0.19361 |
| Chitin 6h | 442.536 | 37.8927 | 2.51547 | 0.146957 |
| Epi 6h | 391.731 | 55.3317 | 2.07523 | 0.235511 |
| SA 6h | 377.312 | 27.5195 | 2.31301 | 0.329742 |
| Me-JA 6h | 424.77 | 56.476 | 2.55098 | 0.148957 |
Source Transcript PGSC0003DMT400015269 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G71010.1 | +2 | 0.0 | 772 | 421/774 (54%) | FORMS APLOID AND BINUCLEATE CELLS 1C | chr1:26782839-26788712 FORWARD LENGTH=1648 |
