Probe CUST_38813_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38813_PI426222305 | JHI_St_60k_v1 | DMT400015298 | CCTTTTTGGTGTTAACTTGTAGAGTTTATCTAACTCTAGTTCTACCCCAAGATCTTGTAT |
All Microarray Probes Designed to Gene DMG401005966
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38813_PI426222305 | JHI_St_60k_v1 | DMT400015298 | CCTTTTTGGTGTTAACTTGTAGAGTTTATCTAACTCTAGTTCTACCCCAAGATCTTGTAT |
| CUST_38771_PI426222305 | JHI_St_60k_v1 | DMT400015292 | AATGTGGCGCTTATCGCGATAAATATATGCACTAATTGGGATAAGGAGAAATCGCCAGCT |
Microarray Signals from CUST_38813_PI426222305
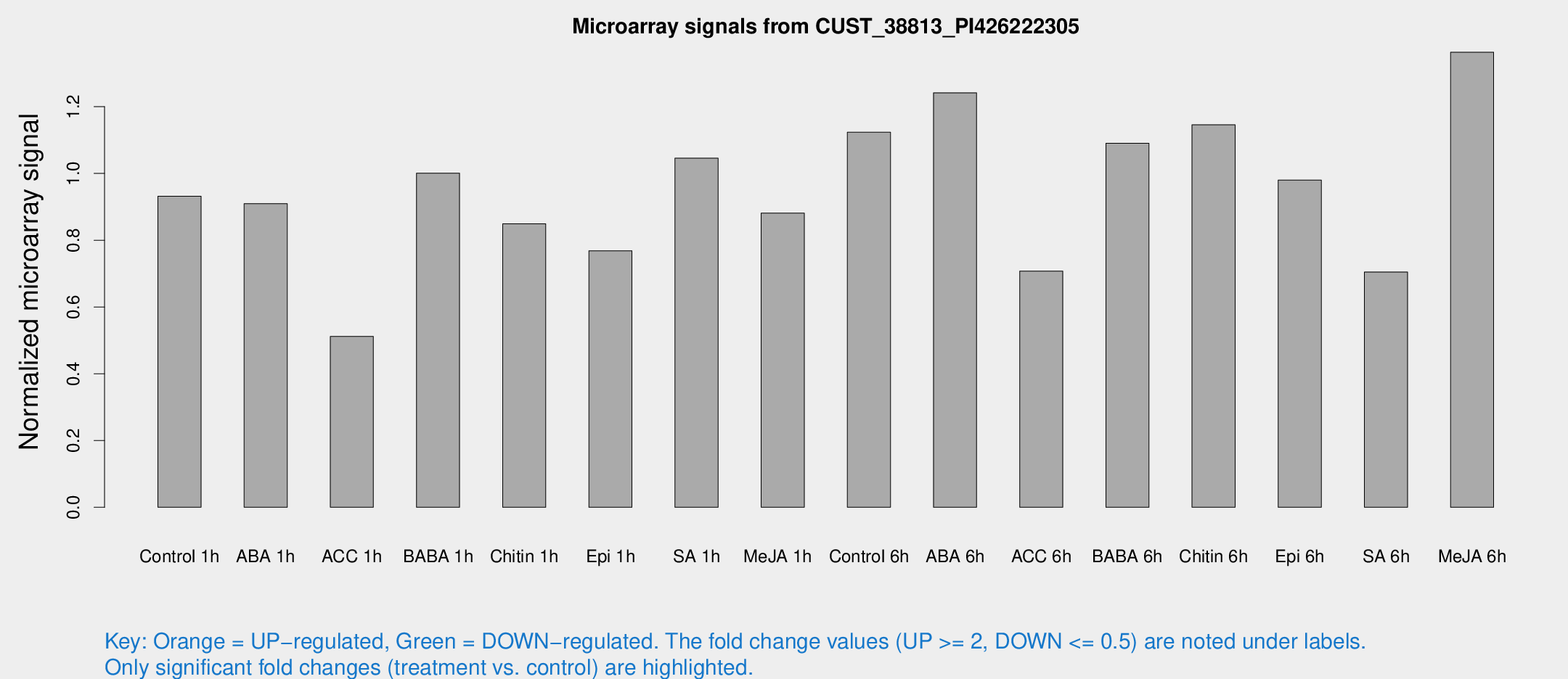
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 62.8369 | 5.01293 | 0.931868 | 0.0741804 |
| ABA 1h | 55.7434 | 9.05355 | 0.909586 | 0.132643 |
| ACC 1h | 46.0954 | 18.321 | 0.511957 | 0.313442 |
| BABA 1h | 65.8884 | 7.25811 | 1.00079 | 0.0845755 |
| Chitin 1h | 52.5976 | 8.03824 | 0.849244 | 0.19131 |
| Epi 1h | 51.2858 | 19.2563 | 0.768457 | 0.308504 |
| SA 1h | 75.2013 | 13.6114 | 1.04561 | 0.287192 |
| Me-JA 1h | 60.2419 | 29.2476 | 0.881087 | 0.365783 |
| Control 6h | 78.9207 | 14.594 | 1.12336 | 0.122594 |
| ABA 6h | 99.239 | 31.2315 | 1.24157 | 0.374628 |
| ACC 6h | 55.1639 | 5.714 | 0.70752 | 0.168637 |
| BABA 6h | 84.6556 | 13.3456 | 1.09043 | 0.152451 |
| Chitin 6h | 82.4616 | 6.37179 | 1.14571 | 0.0887017 |
| Epi 6h | 77.0003 | 14.6221 | 0.980182 | 0.181739 |
| SA 6h | 54.9524 | 17.7037 | 0.704821 | 0.426572 |
| Me-JA 6h | 93.928 | 14.4543 | 1.36308 | 0.150727 |
Source Transcript PGSC0003DMT400015298 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G12700.1 | +2 | 4e-115 | 382 | 190/509 (37%) | ATP binding;nucleic acid binding;helicases | chr1:4323722-4326227 REVERSE LENGTH=735 |
