Probe CUST_38771_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38771_PI426222305 | JHI_St_60k_v1 | DMT400015292 | AATGTGGCGCTTATCGCGATAAATATATGCACTAATTGGGATAAGGAGAAATCGCCAGCT |
All Microarray Probes Designed to Gene DMG401005966
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38813_PI426222305 | JHI_St_60k_v1 | DMT400015298 | CCTTTTTGGTGTTAACTTGTAGAGTTTATCTAACTCTAGTTCTACCCCAAGATCTTGTAT |
| CUST_38771_PI426222305 | JHI_St_60k_v1 | DMT400015292 | AATGTGGCGCTTATCGCGATAAATATATGCACTAATTGGGATAAGGAGAAATCGCCAGCT |
Microarray Signals from CUST_38771_PI426222305
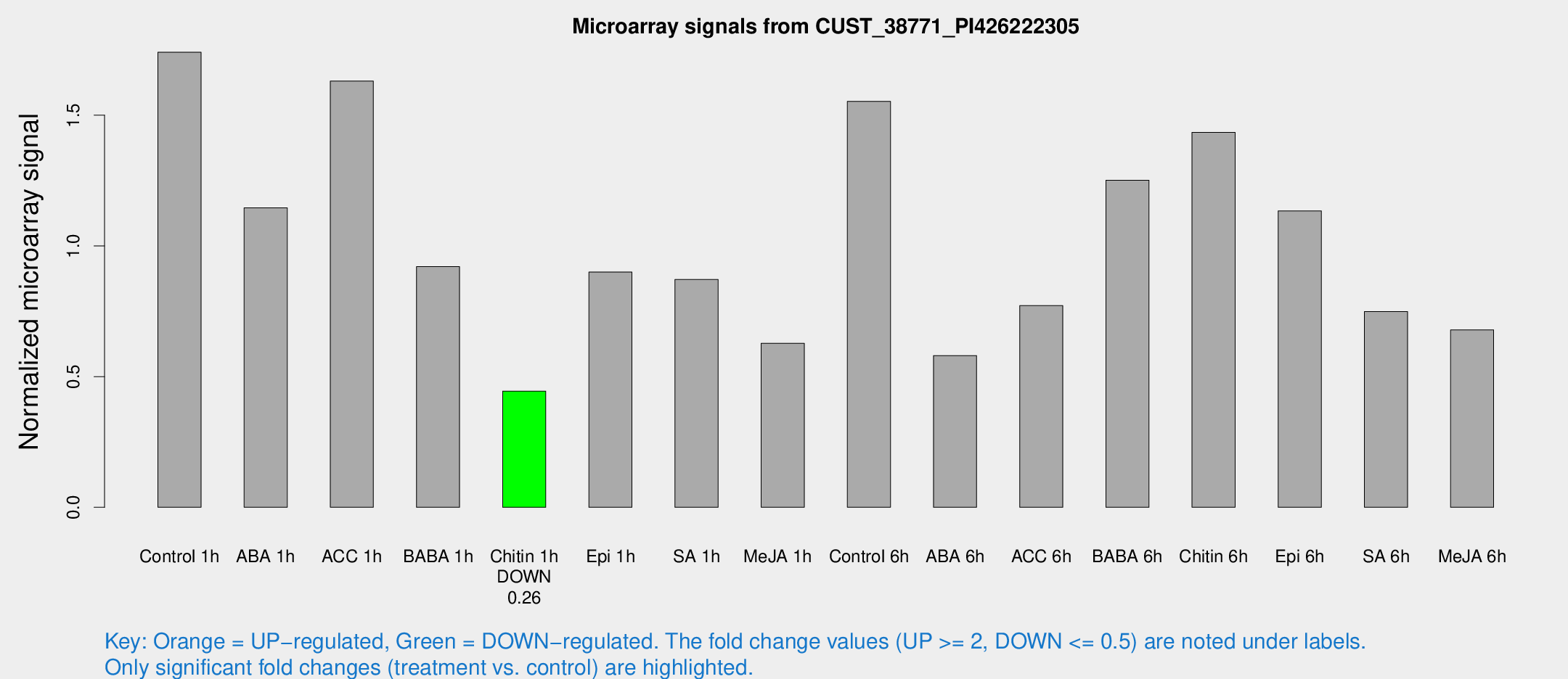
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 31.9327 | 4.59679 | 1.741 | 0.27706 |
| ABA 1h | 18.3528 | 3.30064 | 1.14598 | 0.212439 |
| ACC 1h | 30.0486 | 4.42869 | 1.63091 | 0.243747 |
| BABA 1h | 16.8233 | 4.00644 | 0.920794 | 0.238187 |
| Chitin 1h | 7.5993 | 3.24569 | 0.444575 | 0.212306 |
| Epi 1h | 14.3797 | 3.26551 | 0.899913 | 0.224261 |
| SA 1h | 16.1393 | 3.42553 | 0.871584 | 0.187091 |
| Me-JA 1h | 10.3522 | 3.61205 | 0.627596 | 0.286853 |
| Control 6h | 34.9342 | 16.1996 | 1.553 | 0.734019 |
| ABA 6h | 12.1736 | 3.52752 | 0.580444 | 0.211502 |
| ACC 6h | 16.0961 | 4.09354 | 0.771676 | 0.205879 |
| BABA 6h | 27.8449 | 9.19919 | 1.25142 | 0.399577 |
| Chitin 6h | 28.0468 | 4.40057 | 1.43456 | 0.218865 |
| Epi 6h | 23.2518 | 4.17971 | 1.13358 | 0.207394 |
| SA 6h | 19.1677 | 11.765 | 0.748788 | 0.498713 |
| Me-JA 6h | 13.7923 | 4.25093 | 0.678709 | 0.235862 |
Source Transcript PGSC0003DMT400015292 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G52450.1 | +1 | 1e-112 | 342 | 166/277 (60%) | MATE efflux family protein | chr5:21289042-21291749 REVERSE LENGTH=486 |
