Probe CUST_38383_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38383_PI426222305 | JHI_St_60k_v1 | DMT400074356 | CTGGTCTCTAAGATATCTCCACCATTTCCTCTGTATAATTCGTCACTATCTCTTTTTCGG |
All Microarray Probes Designed to Gene DMG401028902
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38383_PI426222305 | JHI_St_60k_v1 | DMT400074356 | CTGGTCTCTAAGATATCTCCACCATTTCCTCTGTATAATTCGTCACTATCTCTTTTTCGG |
| CUST_38444_PI426222305 | JHI_St_60k_v1 | DMT400074357 | CTGGTCTCTAAGATATCTCCACCATTTCCTCTGTATAATTCGTCACTATCTCTTTTTCGG |
Microarray Signals from CUST_38383_PI426222305
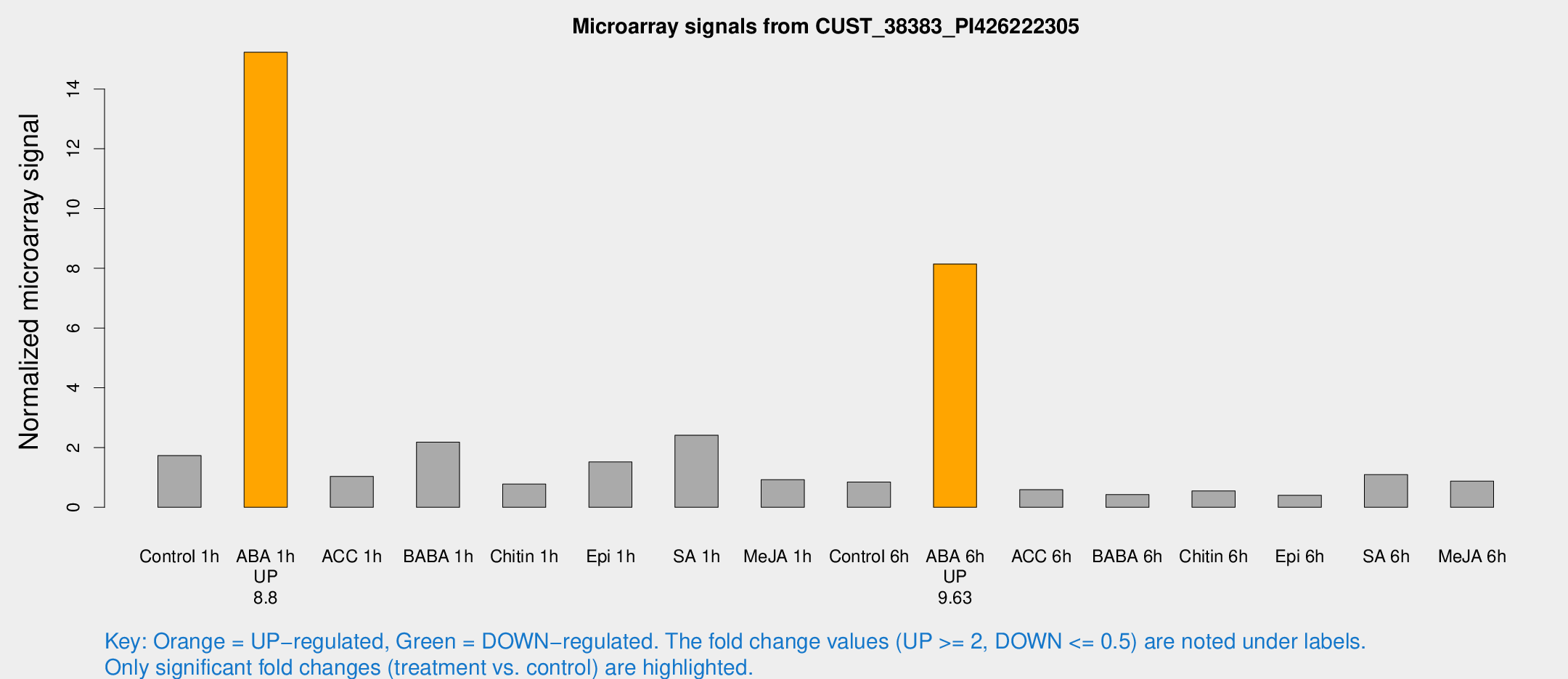
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 33.0517 | 7.51583 | 1.7307 | 0.509035 |
| ABA 1h | 241.477 | 14.2846 | 15.2318 | 1.22941 |
| ACC 1h | 22.6818 | 7.83216 | 1.03665 | 0.437167 |
| BABA 1h | 58.4693 | 38.5935 | 2.17857 | 1.70975 |
| Chitin 1h | 14.9846 | 5.67587 | 0.776152 | 0.39035 |
| Epi 1h | 27.8299 | 10.4879 | 1.51695 | 0.742389 |
| SA 1h | 61.7897 | 34.3012 | 2.41438 | 2.18918 |
| Me-JA 1h | 14.84 | 4.78777 | 0.92529 | 0.383285 |
| Control 6h | 16.217 | 4.30072 | 0.845758 | 0.242268 |
| ABA 6h | 158.054 | 21.3167 | 8.14499 | 0.892859 |
| ACC 6h | 12.1226 | 3.97267 | 0.589031 | 0.187755 |
| BABA 6h | 8.9568 | 3.56515 | 0.425478 | 0.188963 |
| Chitin 6h | 11.4711 | 3.57046 | 0.551244 | 0.211072 |
| Epi 6h | 8.12554 | 3.73237 | 0.40089 | 0.187287 |
| SA 6h | 20.9385 | 5.73043 | 1.09369 | 0.230711 |
| Me-JA 6h | 16.6873 | 4.09336 | 0.875372 | 0.202926 |
Source Transcript PGSC0003DMT400074356 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G24080.1 | +1 | 0.0 | 660 | 331/470 (70%) | Protein kinase superfamily protein | chr5:8139334-8141014 REVERSE LENGTH=470 |
