Probe CUST_37647_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37647_PI426222305 | JHI_St_60k_v1 | DMT400049665 | CATGATTATTGAGGTGGAAAGGGCATTTATTCTTTGGTTGGCGACTAAAATAGCAAAAAA |
All Microarray Probes Designed to Gene DMG400019294
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37647_PI426222305 | JHI_St_60k_v1 | DMT400049665 | CATGATTATTGAGGTGGAAAGGGCATTTATTCTTTGGTTGGCGACTAAAATAGCAAAAAA |
| CUST_37640_PI426222305 | JHI_St_60k_v1 | DMT400049666 | CATGATTATTGAGGTGGAAAGGGCATTTATTCTTTGGTTGGCGACTAAAATAGCAAAAAA |
Microarray Signals from CUST_37647_PI426222305
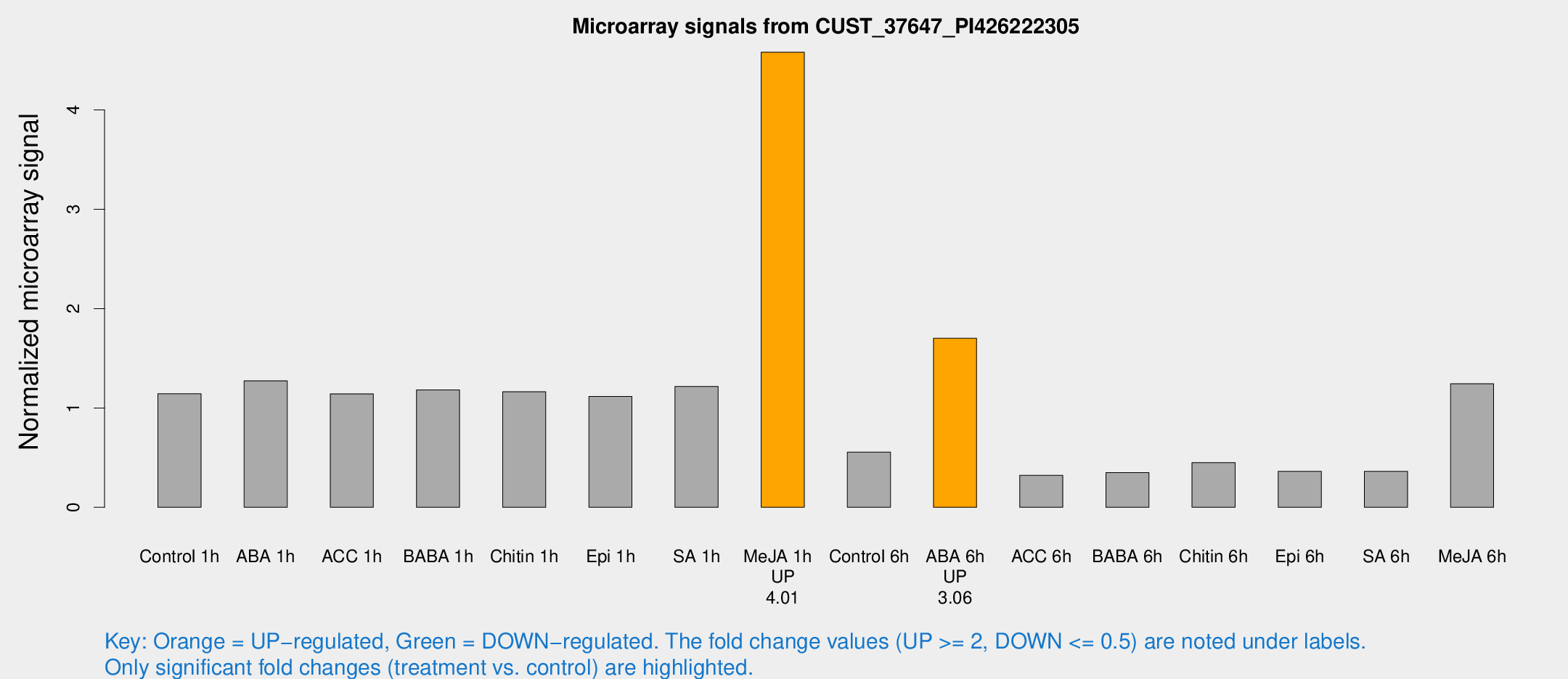
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5803.69 | 872.933 | 1.14309 | 0.143718 |
| ABA 1h | 5767.59 | 998.372 | 1.27332 | 0.138744 |
| ACC 1h | 6353.47 | 1978.68 | 1.14195 | 0.319511 |
| BABA 1h | 5869.45 | 1056.84 | 1.18186 | 0.114481 |
| Chitin 1h | 5197.39 | 300.43 | 1.16295 | 0.129948 |
| Epi 1h | 4862.47 | 585.582 | 1.11535 | 0.134845 |
| SA 1h | 6408.85 | 1169.42 | 1.21719 | 0.204987 |
| Me-JA 1h | 18787.4 | 2007.21 | 4.58106 | 0.264489 |
| Control 6h | 2830.51 | 441.596 | 0.555985 | 0.0536045 |
| ABA 6h | 9260.4 | 1618.33 | 1.70135 | 0.206564 |
| ACC 6h | 1842.03 | 107.491 | 0.322361 | 0.062415 |
| BABA 6h | 1954.29 | 150.906 | 0.349497 | 0.0414956 |
| Chitin 6h | 2473.39 | 445.901 | 0.450391 | 0.109499 |
| Epi 6h | 2052.38 | 236.368 | 0.361381 | 0.0720508 |
| SA 6h | 1867.19 | 381.009 | 0.361844 | 0.0423845 |
| Me-JA 6h | 6518.52 | 1447.16 | 1.24306 | 0.340426 |
Source Transcript PGSC0003DMT400049665 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27410.3 | +3 | 4e-119 | 356 | 185/277 (67%) | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | chr4:13707928-13709013 REVERSE LENGTH=314 |
