Probe CUST_37535_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37535_PI426222305 | JHI_St_60k_v1 | DMT400014217 | GTCATCTGAGTGCCGTTCATATCGAGATTCTAGATACTTTTGTTCTTCTTGTGAAAATTT |
All Microarray Probes Designed to Gene DMG400005573
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37554_PI426222305 | JHI_St_60k_v1 | DMT400014216 | ATGAATATGCAATTGGGCAACTGAAGGAATATGATGATGGCGGATGTGCAGATGGCTGAG |
| CUST_37535_PI426222305 | JHI_St_60k_v1 | DMT400014217 | GTCATCTGAGTGCCGTTCATATCGAGATTCTAGATACTTTTGTTCTTCTTGTGAAAATTT |
Microarray Signals from CUST_37535_PI426222305
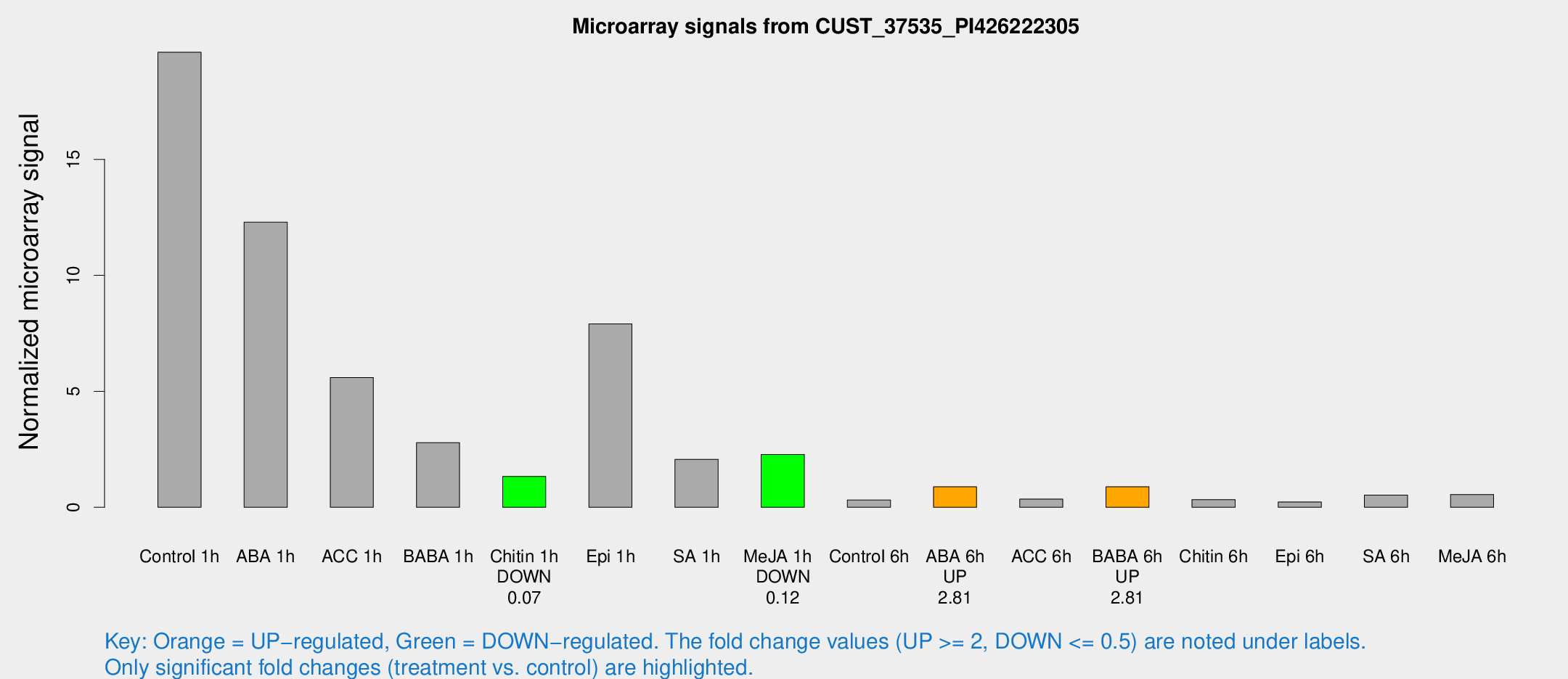
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2824.26 | 481.937 | 19.6193 | 3.1722 |
| ABA 1h | 3051.52 | 2279.95 | 12.2917 | 22.3059 |
| ACC 1h | 1665.4 | 1141.73 | 5.59669 | 10.3365 |
| BABA 1h | 631.131 | 432.819 | 2.78726 | 2.52558 |
| Chitin 1h | 278.974 | 188.633 | 1.33182 | 1.2832 |
| Epi 1h | 1230.39 | 513.932 | 7.90633 | 5.88065 |
| SA 1h | 430.221 | 205.424 | 2.07045 | 1.86771 |
| Me-JA 1h | 262.114 | 23.7683 | 2.27498 | 0.134568 |
| Control 6h | 45.0463 | 6.74487 | 0.314301 | 0.0311264 |
| ABA 6h | 140.319 | 36.5611 | 0.883862 | 0.223412 |
| ACC 6h | 57.2451 | 5.32258 | 0.354705 | 0.0714099 |
| BABA 6h | 141.204 | 19.0873 | 0.884729 | 0.158726 |
| Chitin 6h | 50.8736 | 10.0113 | 0.328782 | 0.0722995 |
| Epi 6h | 36.7954 | 4.51402 | 0.23275 | 0.0285208 |
| SA 6h | 76.6906 | 17.7561 | 0.523629 | 0.0754709 |
| Me-JA 6h | 77.3831 | 5.62323 | 0.552724 | 0.0434336 |
Source Transcript PGSC0003DMT400014217 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G52640.1 | +1 | 0.0 | 409 | 242/275 (88%) | heat shock protein 90.1 | chr5:21352542-21355147 FORWARD LENGTH=705 |
