Probe CUST_37200_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37200_PI426222305 | JHI_St_60k_v1 | DMT400082472 | TTGCAGCATCAATTATAACTGAAATTGATCTTTACAAGTTTGATCCATGGCAGTTGCCTG |
All Microarray Probes Designed to Gene DMG400032555
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37192_PI426222305 | JHI_St_60k_v1 | DMT400082471 | TTTGACATATATTGCAGCTTGATGATTGGGTATTGTGTCGAATTTACAACAAGAAAGGCA |
| CUST_37200_PI426222305 | JHI_St_60k_v1 | DMT400082472 | TTGCAGCATCAATTATAACTGAAATTGATCTTTACAAGTTTGATCCATGGCAGTTGCCTG |
Microarray Signals from CUST_37200_PI426222305
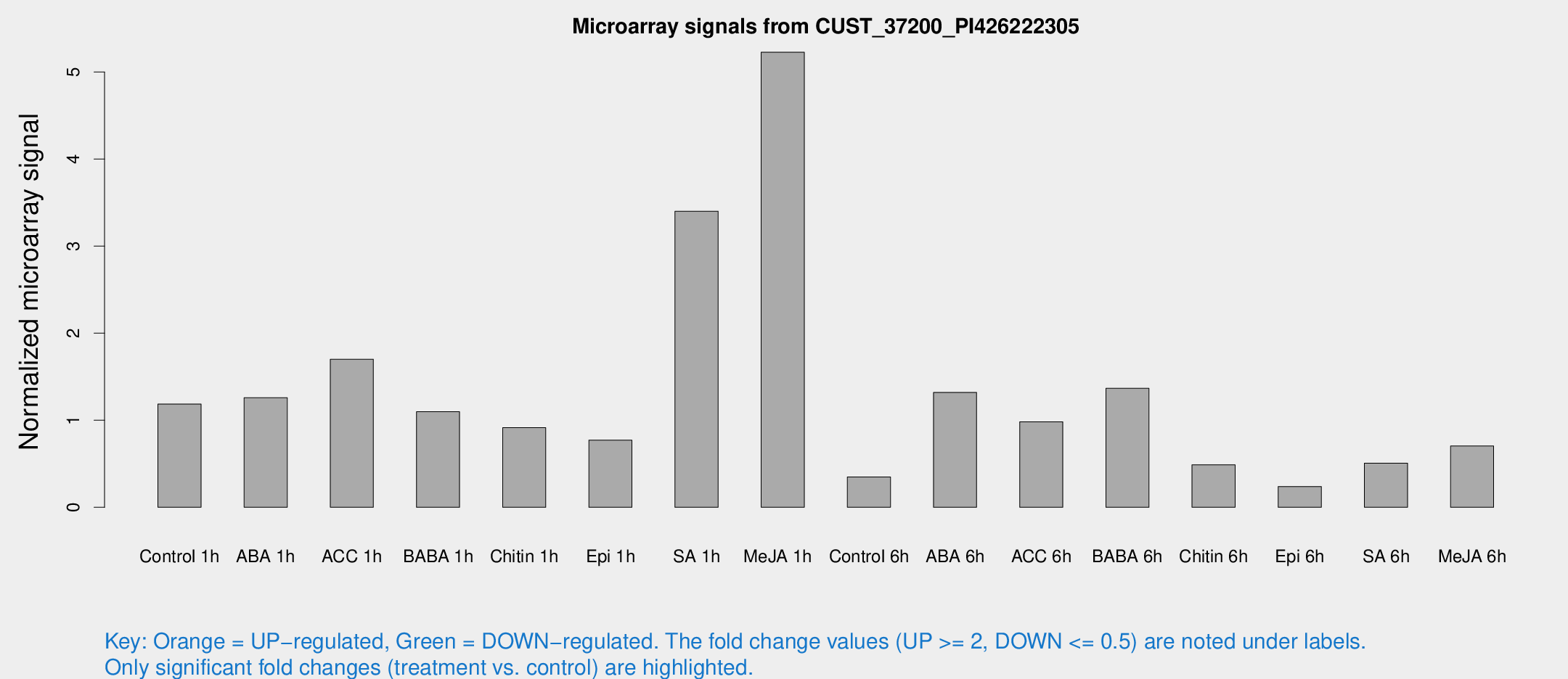
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 761.03 | 228.471 | 1.18613 | 0.274534 |
| ABA 1h | 667.476 | 75.1113 | 1.25857 | 0.072985 |
| ACC 1h | 1263.81 | 492.744 | 1.70063 | 0.79769 |
| BABA 1h | 661.215 | 140.125 | 1.09851 | 0.155274 |
| Chitin 1h | 512.514 | 110.012 | 0.914983 | 0.210888 |
| Epi 1h | 426.302 | 104.783 | 0.7712 | 0.224115 |
| SA 1h | 2232.91 | 642.499 | 3.40178 | 0.75061 |
| Me-JA 1h | 2702.66 | 705.9 | 5.22795 | 0.931923 |
| Control 6h | 276.045 | 117.55 | 0.34819 | 0.217619 |
| ABA 6h | 1044.23 | 393.924 | 1.31962 | 0.773122 |
| ACC 6h | 695.613 | 135.129 | 0.982497 | 0.12698 |
| BABA 6h | 999.633 | 328.848 | 1.36664 | 0.394865 |
| Chitin 6h | 329.19 | 78.7067 | 0.488921 | 0.144512 |
| Epi 6h | 175.796 | 57.1885 | 0.238582 | 0.0860798 |
| SA 6h | 304.966 | 50.705 | 0.507667 | 0.0952126 |
| Me-JA 6h | 428.83 | 80.9448 | 0.704182 | 0.0756009 |
Source Transcript PGSC0003DMT400082472 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G01720.1 | +2 | 2e-109 | 328 | 179/304 (59%) | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | chr1:268471-269514 FORWARD LENGTH=289 |
