Probe CUST_37192_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37192_PI426222305 | JHI_St_60k_v1 | DMT400082471 | TTTGACATATATTGCAGCTTGATGATTGGGTATTGTGTCGAATTTACAACAAGAAAGGCA |
All Microarray Probes Designed to Gene DMG400032555
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37192_PI426222305 | JHI_St_60k_v1 | DMT400082471 | TTTGACATATATTGCAGCTTGATGATTGGGTATTGTGTCGAATTTACAACAAGAAAGGCA |
| CUST_37200_PI426222305 | JHI_St_60k_v1 | DMT400082472 | TTGCAGCATCAATTATAACTGAAATTGATCTTTACAAGTTTGATCCATGGCAGTTGCCTG |
Microarray Signals from CUST_37192_PI426222305
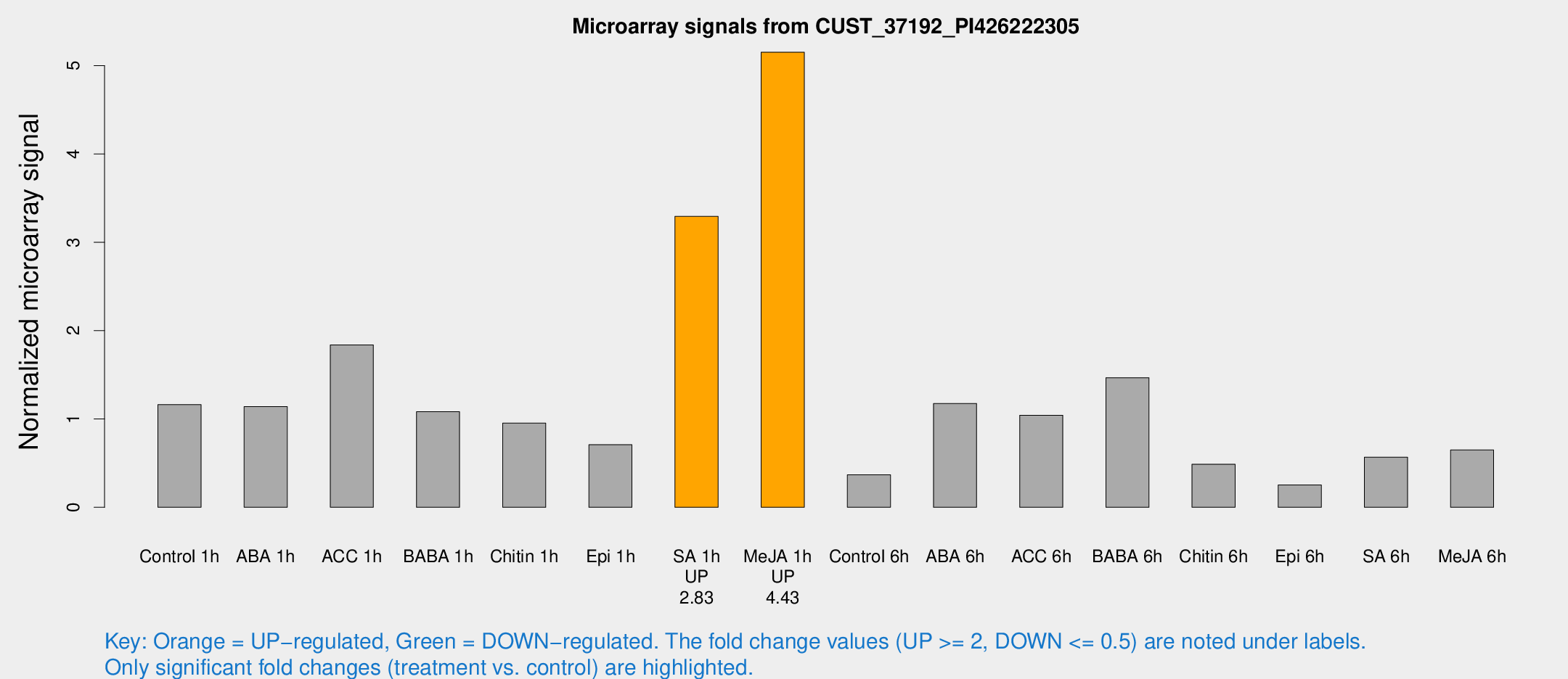
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1263.42 | 301.796 | 1.16171 | 0.196579 |
| ABA 1h | 1051.09 | 100.646 | 1.1393 | 0.0658963 |
| ACC 1h | 2154.62 | 696.483 | 1.83709 | 0.513989 |
| BABA 1h | 1125.37 | 225.549 | 1.08144 | 0.133034 |
| Chitin 1h | 906.742 | 134.706 | 0.952982 | 0.156358 |
| Epi 1h | 681.927 | 163.414 | 0.708322 | 0.204903 |
| SA 1h | 3765.28 | 1041.82 | 3.29311 | 0.716408 |
| Me-JA 1h | 4616.42 | 1150.03 | 5.15088 | 0.838227 |
| Control 6h | 481.857 | 212.901 | 0.368241 | 0.175646 |
| ABA 6h | 1516.64 | 509.96 | 1.17487 | 0.510839 |
| ACC 6h | 1279.38 | 222.142 | 1.04233 | 0.16276 |
| BABA 6h | 1819.37 | 506.974 | 1.46693 | 0.353616 |
| Chitin 6h | 553.326 | 99.5352 | 0.48791 | 0.0983502 |
| Epi 6h | 308.446 | 66.4132 | 0.253934 | 0.0543941 |
| SA 6h | 597.218 | 99.3324 | 0.567322 | 0.0433553 |
| Me-JA 6h | 675.625 | 78.7422 | 0.648441 | 0.0376412 |
Source Transcript PGSC0003DMT400082471 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G08790.1 | +1 | 8e-53 | 181 | 84/91 (92%) | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | chr5:2859113-2860144 REVERSE LENGTH=283 |
