Probe CUST_36801_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36801_PI426222305 | JHI_St_60k_v1 | DMT400004181 | CACGTTTAGATTATTGATTAATGGTGGACTGGAGAGATTTGTTCATCTTTGGCACGTTAT |
All Microarray Probes Designed to Gene DMG400001662
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36801_PI426222305 | JHI_St_60k_v1 | DMT400004181 | CACGTTTAGATTATTGATTAATGGTGGACTGGAGAGATTTGTTCATCTTTGGCACGTTAT |
Microarray Signals from CUST_36801_PI426222305
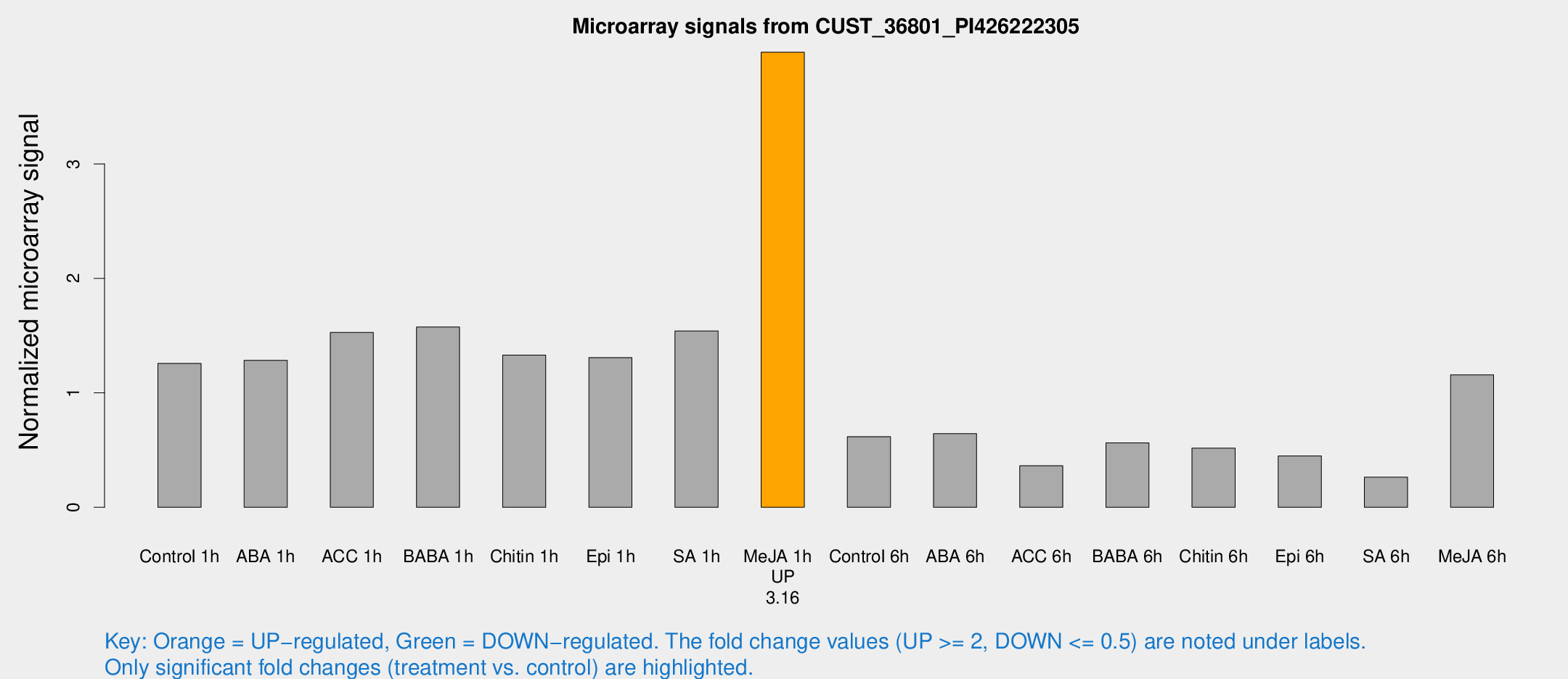
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 207.545 | 15.6665 | 1.25714 | 0.103605 |
| ABA 1h | 205.553 | 67.4313 | 1.28442 | 0.330393 |
| ACC 1h | 267.117 | 48.8118 | 1.52864 | 0.198703 |
| BABA 1h | 250.472 | 17.1649 | 1.57556 | 0.0930869 |
| Chitin 1h | 200.342 | 29.7599 | 1.32991 | 0.219286 |
| Epi 1h | 187.531 | 18.0395 | 1.30819 | 0.128439 |
| SA 1h | 273.634 | 63.89 | 1.54155 | 0.337859 |
| Me-JA 1h | 533.391 | 30.9753 | 3.97663 | 0.281059 |
| Control 6h | 107.123 | 22.3843 | 0.617176 | 0.0921208 |
| ABA 6h | 114.749 | 17.6043 | 0.643316 | 0.061222 |
| ACC 6h | 71.18 | 14.0794 | 0.363014 | 0.049354 |
| BABA 6h | 103.538 | 6.89183 | 0.563014 | 0.0374225 |
| Chitin 6h | 93.4266 | 17.0703 | 0.517077 | 0.0909484 |
| Epi 6h | 86.1642 | 16.9095 | 0.448605 | 0.10366 |
| SA 6h | 53.8386 | 20.5966 | 0.262804 | 0.126717 |
| Me-JA 6h | 210.706 | 70.9109 | 1.15709 | 0.295142 |
Source Transcript PGSC0003DMT400004181 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G34710.1 | +2 | 0.0 | 911 | 503/694 (72%) | arginine decarboxylase 2 | chr4:16560315-16562450 REVERSE LENGTH=711 |
