Probe CUST_36742_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36742_PI426222305 | JHI_St_60k_v1 | DMT400087067 | GACAAGGGCATCACCATTCCCTGTGAAGAATTCATTTTCCAATCACTCACTTCCATGATC |
All Microarray Probes Designed to Gene DMG400036638
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36742_PI426222305 | JHI_St_60k_v1 | DMT400087067 | GACAAGGGCATCACCATTCCCTGTGAAGAATTCATTTTCCAATCACTCACTTCCATGATC |
Microarray Signals from CUST_36742_PI426222305
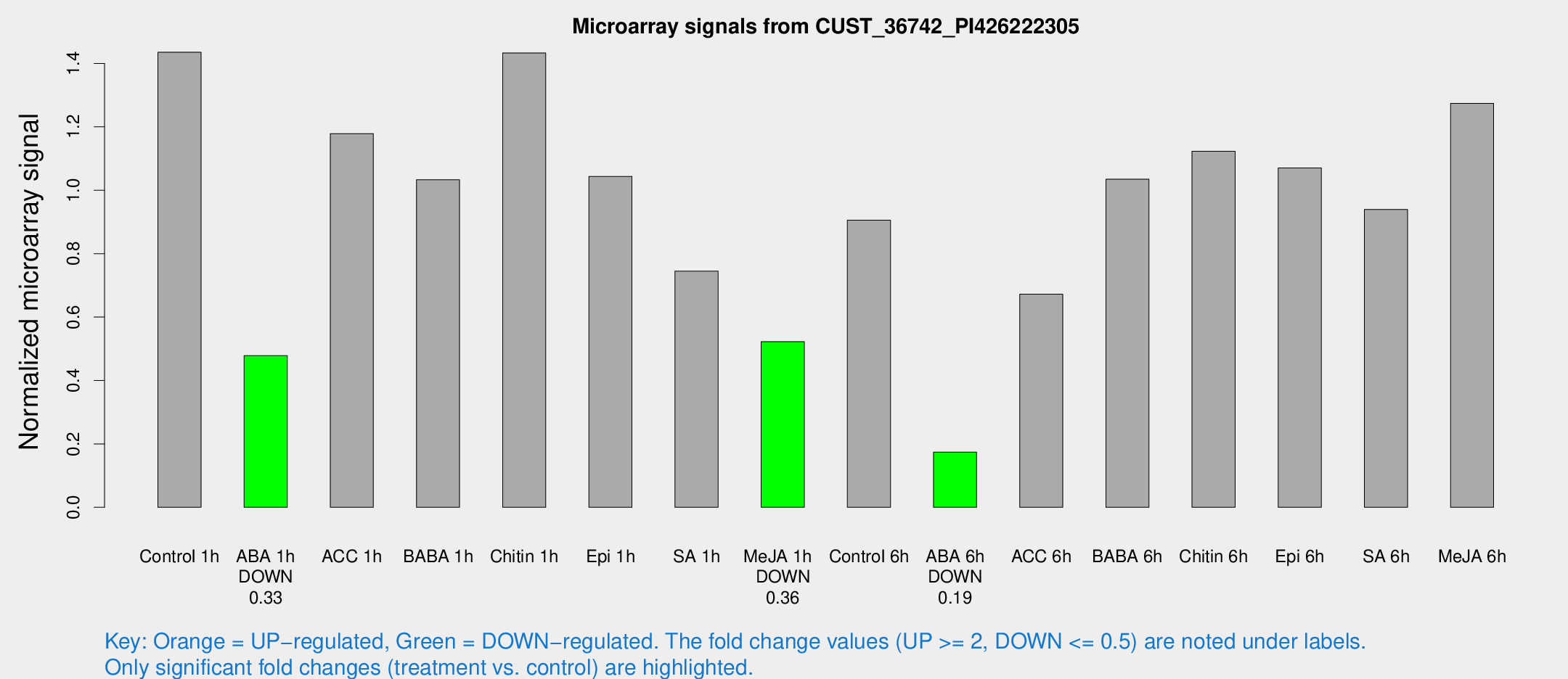
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 558.189 | 90.7359 | 1.43522 | 0.196888 |
| ABA 1h | 161.194 | 15.6422 | 0.478263 | 0.0291578 |
| ACC 1h | 457.906 | 26.7718 | 1.17857 | 0.0965513 |
| BABA 1h | 390.268 | 69.8306 | 1.03302 | 0.0992864 |
| Chitin 1h | 498.325 | 80.2734 | 1.43275 | 0.226099 |
| Epi 1h | 346.048 | 43.4589 | 1.04362 | 0.101964 |
| SA 1h | 300.455 | 57.8196 | 0.744613 | 0.107097 |
| Me-JA 1h | 162.007 | 13.7164 | 0.522151 | 0.0319507 |
| Control 6h | 357.924 | 68.2113 | 0.905789 | 0.112984 |
| ABA 6h | 70.9612 | 9.42605 | 0.173931 | 0.0156233 |
| ACC 6h | 302.322 | 59.8482 | 0.672252 | 0.0449415 |
| BABA 6h | 465.633 | 110.543 | 1.03478 | 0.245534 |
| Chitin 6h | 454.986 | 42.7226 | 1.12274 | 0.0685654 |
| Epi 6h | 465.183 | 65.2933 | 1.07019 | 0.224601 |
| SA 6h | 355.31 | 38.717 | 0.939606 | 0.208704 |
| Me-JA 6h | 497.523 | 89.7813 | 1.27389 | 0.173135 |
Source Transcript PGSC0003DMT400087067 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G34760.1 | +1 | 2e-45 | 145 | 75/109 (69%) | SAUR-like auxin-responsive protein family | chr4:16582471-16582794 REVERSE LENGTH=107 |
