Probe CUST_36553_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36553_PI426222305 | JHI_St_60k_v1 | DMT400064551 | TGTAATGTAAGAAGATGGTGTTTTGGATCTCTTCTCAAAATGTTTTCTAGCTCCATGATG |
All Microarray Probes Designed to Gene DMG401025083
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36553_PI426222305 | JHI_St_60k_v1 | DMT400064551 | TGTAATGTAAGAAGATGGTGTTTTGGATCTCTTCTCAAAATGTTTTCTAGCTCCATGATG |
| CUST_36514_PI426222305 | JHI_St_60k_v1 | DMT400064554 | AAGTTGTGTCTTGCTCTGATATTACTGCCATTGCTGCTAGGGACTCTGTTGTTTTGACGG |
Microarray Signals from CUST_36553_PI426222305
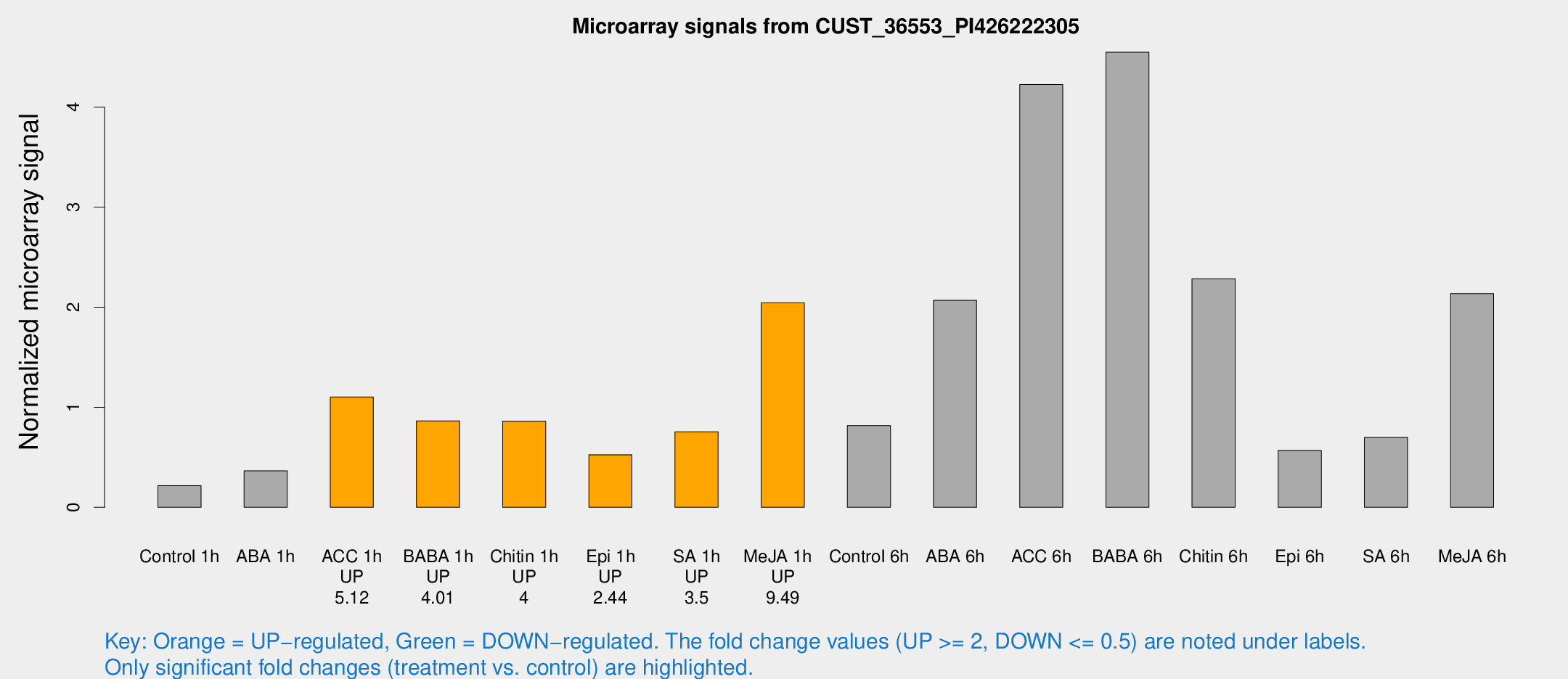
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 93.222 | 21.912 | 0.215454 | 0.0480489 |
| ABA 1h | 153.026 | 63.7951 | 0.364762 | 0.127112 |
| ACC 1h | 517.613 | 183.847 | 1.10272 | 0.332323 |
| BABA 1h | 337.942 | 22.1123 | 0.863959 | 0.137623 |
| Chitin 1h | 361.837 | 131.58 | 0.861313 | 0.337475 |
| Epi 1h | 186.874 | 23.7573 | 0.524911 | 0.0690389 |
| SA 1h | 343.38 | 90.0054 | 0.754695 | 0.217688 |
| Me-JA 1h | 680.802 | 65.377 | 2.04376 | 0.118469 |
| Control 6h | 608.903 | 300.891 | 0.816872 | 1.42023 |
| ABA 6h | 1282.56 | 752.552 | 2.06893 | 1.86803 |
| ACC 6h | 2125.32 | 601.525 | 4.22554 | 0.848148 |
| BABA 6h | 2232.51 | 672.363 | 4.54859 | 1.19207 |
| Chitin 6h | 1121.99 | 425.11 | 2.28487 | 0.918139 |
| Epi 6h | 299.108 | 94.9478 | 0.567999 | 0.320181 |
| SA 6h | 328.534 | 138.593 | 0.698431 | 0.254398 |
| Me-JA 6h | 902.513 | 199.142 | 2.13506 | 0.301016 |
Source Transcript PGSC0003DMT400064551 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G71695.1 | +3 | 2e-79 | 247 | 116/177 (66%) | Peroxidase superfamily protein | chr1:26964359-26966557 FORWARD LENGTH=358 |
