Probe CUST_36368_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36368_PI426222305 | JHI_St_60k_v1 | DMT400079881 | CGCGGTTTGTATGATCCAATGGCAGCTACTTGTACTATAATACAAGTAAAAATAGTAATG |
All Microarray Probes Designed to Gene DMG400031110
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36365_PI426222305 | JHI_St_60k_v1 | DMT400079878 | CGCGGTTTGTATGATCCAATGGCAGCTACTTGTACTATAATACAAGTAAAAATAGTAATG |
| CUST_36368_PI426222305 | JHI_St_60k_v1 | DMT400079881 | CGCGGTTTGTATGATCCAATGGCAGCTACTTGTACTATAATACAAGTAAAAATAGTAATG |
Microarray Signals from CUST_36368_PI426222305
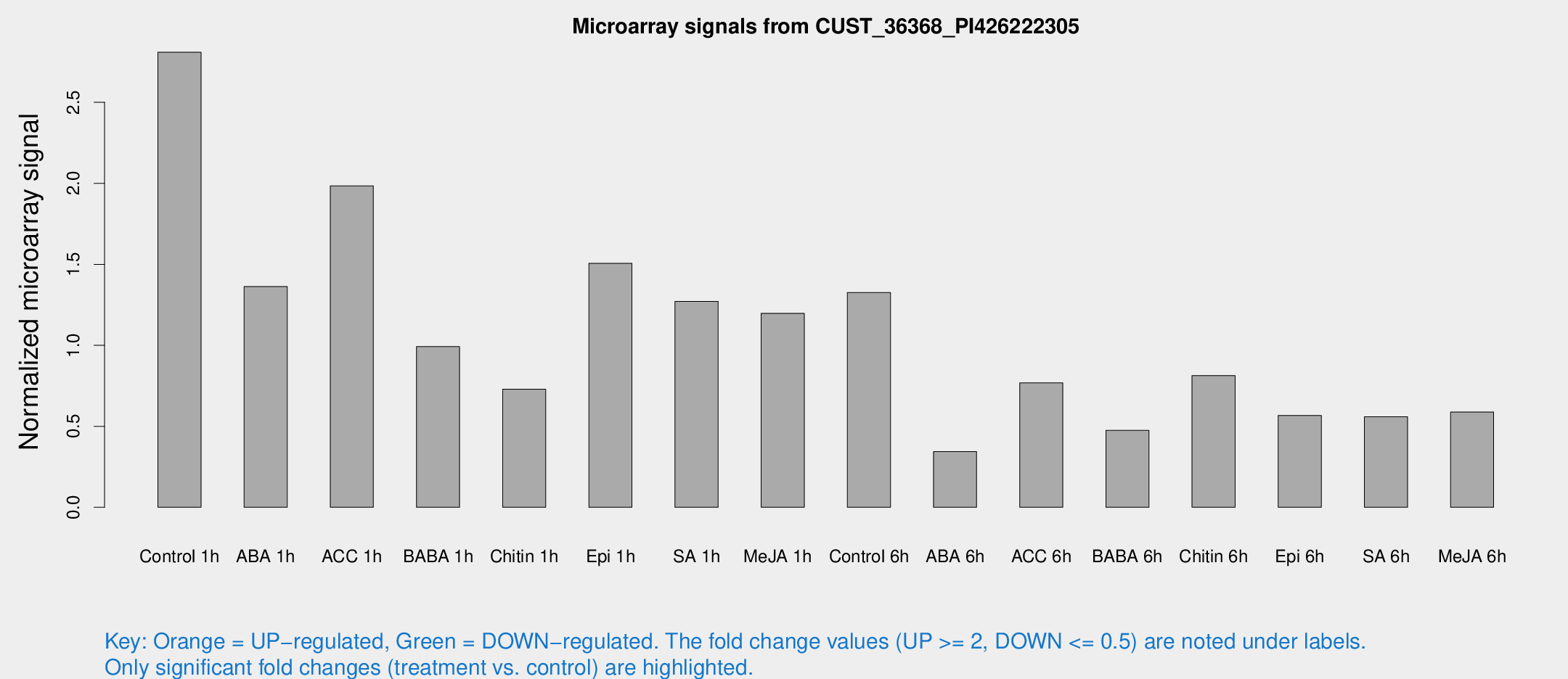
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 45.4965 | 11.1151 | 2.80948 | 0.62777 |
| ABA 1h | 18.4418 | 2.95917 | 1.36242 | 0.22447 |
| ACC 1h | 31.0779 | 3.65814 | 1.98409 | 0.312045 |
| BABA 1h | 16.1549 | 5.71885 | 0.992202 | 0.289769 |
| Chitin 1h | 10.1353 | 2.94005 | 0.728777 | 0.219768 |
| Epi 1h | 19.9706 | 3.09912 | 1.50617 | 0.245623 |
| SA 1h | 19.8323 | 3.10949 | 1.27144 | 0.202655 |
| Me-JA 1h | 16.0855 | 4.98242 | 1.19771 | 0.398388 |
| Control 6h | 21.0262 | 4.76289 | 1.32648 | 0.243031 |
| ABA 6h | 5.51465 | 3.08547 | 0.343973 | 0.192329 |
| ACC 6h | 13.9327 | 3.6782 | 0.767967 | 0.302022 |
| BABA 6h | 8.95261 | 3.35721 | 0.475061 | 0.22352 |
| Chitin 6h | 15.4371 | 5.47808 | 0.813592 | 0.378016 |
| Epi 6h | 10.1411 | 3.46886 | 0.567121 | 0.225263 |
| SA 6h | 9.0219 | 3.17652 | 0.558778 | 0.233005 |
| Me-JA 6h | 9.09734 | 2.98679 | 0.588148 | 0.20862 |
Source Transcript PGSC0003DMT400079881 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G17670.1 | +1 | 1e-69 | 223 | 131/186 (70%) | alpha/beta-Hydrolases superfamily protein | chr5:5821082-5822600 FORWARD LENGTH=309 |
