Probe CUST_35853_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35853_PI426222305 | JHI_St_60k_v1 | DMT400045906 | ACATCTTAAGAGTTGTCGTTTGTGTAGAGTATTTTGGAGGGTCAGCTGACGCAAGCCCTT |
All Microarray Probes Designed to Gene DMG401017804
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35853_PI426222305 | JHI_St_60k_v1 | DMT400045906 | ACATCTTAAGAGTTGTCGTTTGTGTAGAGTATTTTGGAGGGTCAGCTGACGCAAGCCCTT |
| CUST_35844_PI426222305 | JHI_St_60k_v1 | DMT400045908 | GAGGCTAGGGAGTGGACATATTATGAAGATGTGTAAGACGGTTGTAAAAGAATATATGAC |
Microarray Signals from CUST_35853_PI426222305
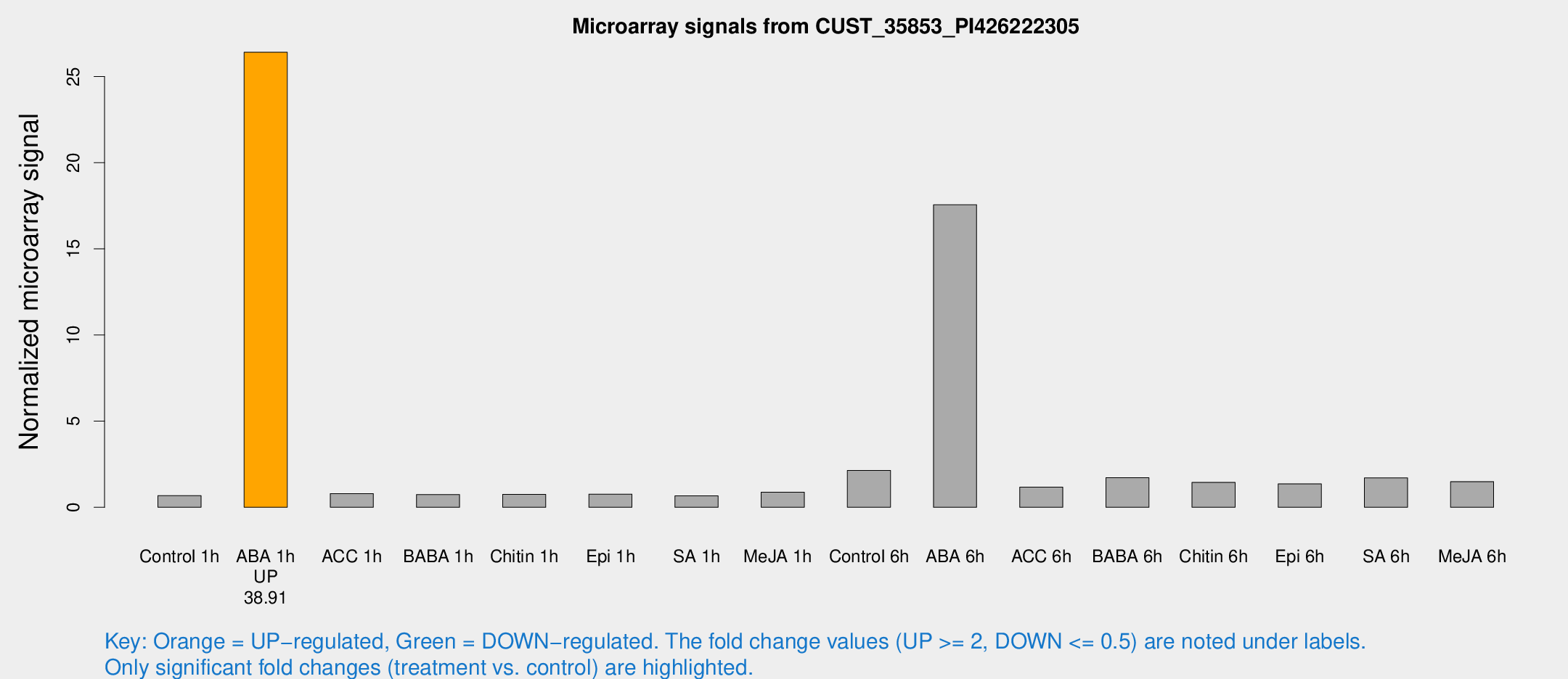
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.91507 | 3.43251 | 0.678759 | 0.393145 |
| ABA 1h | 211.196 | 42.8292 | 26.413 | 5.2513 |
| ACC 1h | 7.33203 | 4.37272 | 0.798071 | 0.462166 |
| BABA 1h | 6.21131 | 3.59948 | 0.740202 | 0.428619 |
| Chitin 1h | 5.8978 | 3.42515 | 0.752189 | 0.435817 |
| Epi 1h | 5.77928 | 3.35203 | 0.766332 | 0.44376 |
| SA 1h | 5.94705 | 3.44508 | 0.665258 | 0.38524 |
| Me-JA 1h | 6.24283 | 3.55162 | 0.877587 | 0.498288 |
| Control 6h | 20.5413 | 6.63739 | 2.13766 | 0.524157 |
| ABA 6h | 174.308 | 41.2917 | 17.5598 | 4.24268 |
| ACC 6h | 11.8472 | 4.437 | 1.17003 | 0.462604 |
| BABA 6h | 17.5533 | 4.02932 | 1.71823 | 0.435586 |
| Chitin 6h | 17.4526 | 9.12917 | 1.44677 | 0.964889 |
| Epi 6h | 15.4474 | 6.15912 | 1.36033 | 0.584724 |
| SA 6h | 17.3434 | 7.44748 | 1.70428 | 1.08454 |
| Me-JA 6h | 17.3007 | 9.61853 | 1.48923 | 0.812749 |
Source Transcript PGSC0003DMT400045906 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G69260.1 | +1 | 1e-64 | 213 | 157/412 (38%) | ABI five binding protein | chr1:26039314-26040570 FORWARD LENGTH=345 |
