Probe CUST_3532_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3532_PI426222305 | JHI_St_60k_v1 | DMT400064255 | GCCAATCAAAGGATCACCACATCCAAGGTATATTTAGAGCTAATTGATCATGAATTTGTT |
All Microarray Probes Designed to Gene DMG400024961
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3532_PI426222305 | JHI_St_60k_v1 | DMT400064255 | GCCAATCAAAGGATCACCACATCCAAGGTATATTTAGAGCTAATTGATCATGAATTTGTT |
| CUST_3535_PI426222305 | JHI_St_60k_v1 | DMT400064256 | ATCAAAGGATCACCACATCCAAGGGGATATTACAAGTGTAGTAGTGTAAGAGGTTGTCCA |
Microarray Signals from CUST_3532_PI426222305
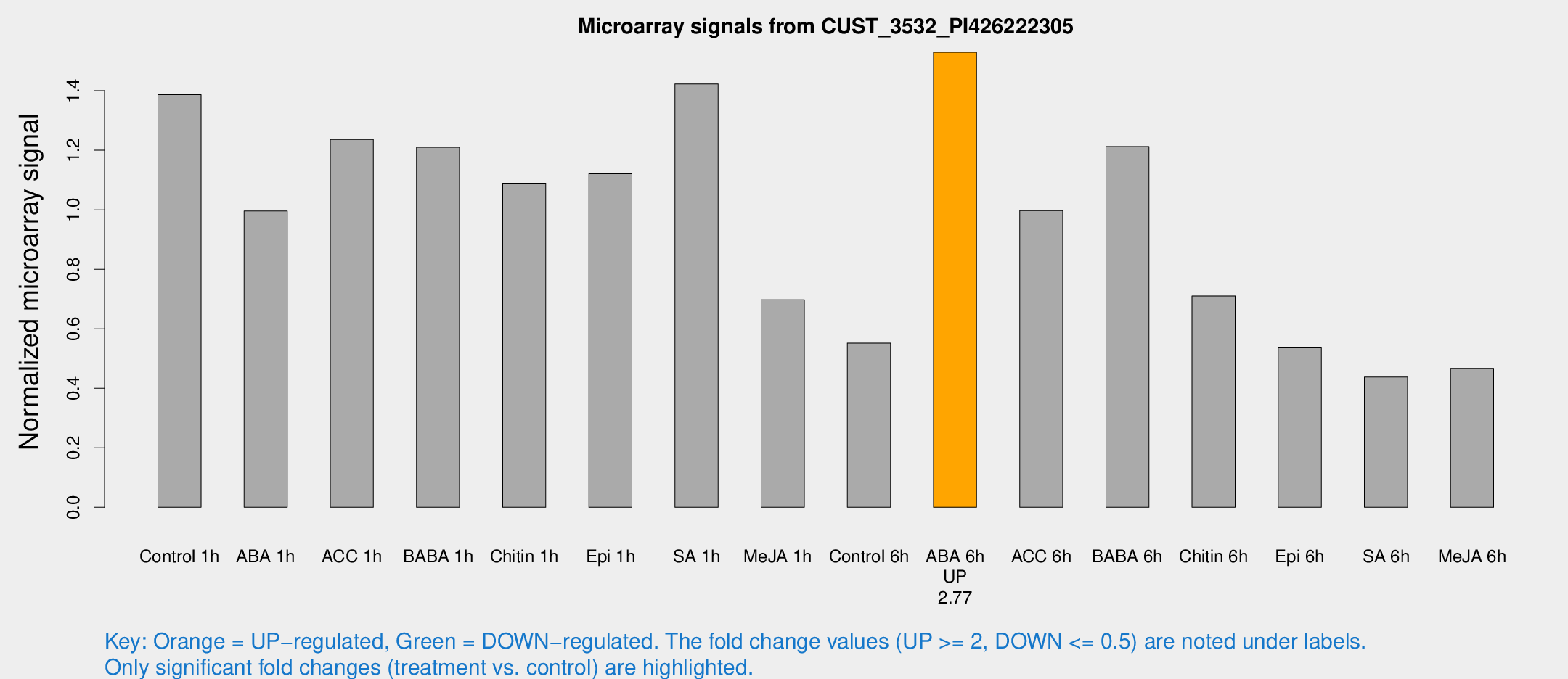
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 687.657 | 113.673 | 1.38671 | 0.132721 |
| ABA 1h | 440.273 | 78.7064 | 0.995966 | 0.116772 |
| ACC 1h | 734.367 | 247.26 | 1.23649 | 0.543381 |
| BABA 1h | 588.021 | 110.51 | 1.21015 | 0.131133 |
| Chitin 1h | 475.557 | 31.5473 | 1.08964 | 0.063421 |
| Epi 1h | 477.421 | 65.9734 | 1.12122 | 0.11906 |
| SA 1h | 717.204 | 84.2822 | 1.42301 | 0.0996912 |
| Me-JA 1h | 279.929 | 38.3265 | 0.697175 | 0.0517655 |
| Control 6h | 299.358 | 85.0641 | 0.552028 | 0.155421 |
| ABA 6h | 806.997 | 127.049 | 1.52941 | 0.173441 |
| ACC 6h | 555.411 | 36.1592 | 0.997382 | 0.143607 |
| BABA 6h | 672.116 | 108.611 | 1.21261 | 0.158344 |
| Chitin 6h | 367.774 | 29.519 | 0.710491 | 0.0621332 |
| Epi 6h | 296.026 | 33.5631 | 0.535963 | 0.0318546 |
| SA 6h | 212.707 | 25.8787 | 0.437717 | 0.0266373 |
| Me-JA 6h | 233.289 | 46.6528 | 0.467268 | 0.0579635 |
Source Transcript PGSC0003DMT400064255 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G23320.1 | +2 | 2e-52 | 139 | 122/293 (42%) | WRKY DNA-binding protein 15 | chr2:9924998-9926154 FORWARD LENGTH=317 |
