Probe CUST_34248_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34248_PI426222305 | JHI_St_60k_v1 | DMT400044819 | CTTGTTGTGTTACATCCTGGGCTATATTTTGCTAATGACTAATAATATACCCTACTCTTC |
All Microarray Probes Designed to Gene DMG400017394
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34248_PI426222305 | JHI_St_60k_v1 | DMT400044819 | CTTGTTGTGTTACATCCTGGGCTATATTTTGCTAATGACTAATAATATACCCTACTCTTC |
| CUST_34231_PI426222305 | JHI_St_60k_v1 | DMT400044818 | CTTGTTGTGTTACATCCTGGGCTATATTTTGCTAATGACTAATAATATACCCTACTCTTC |
Microarray Signals from CUST_34248_PI426222305
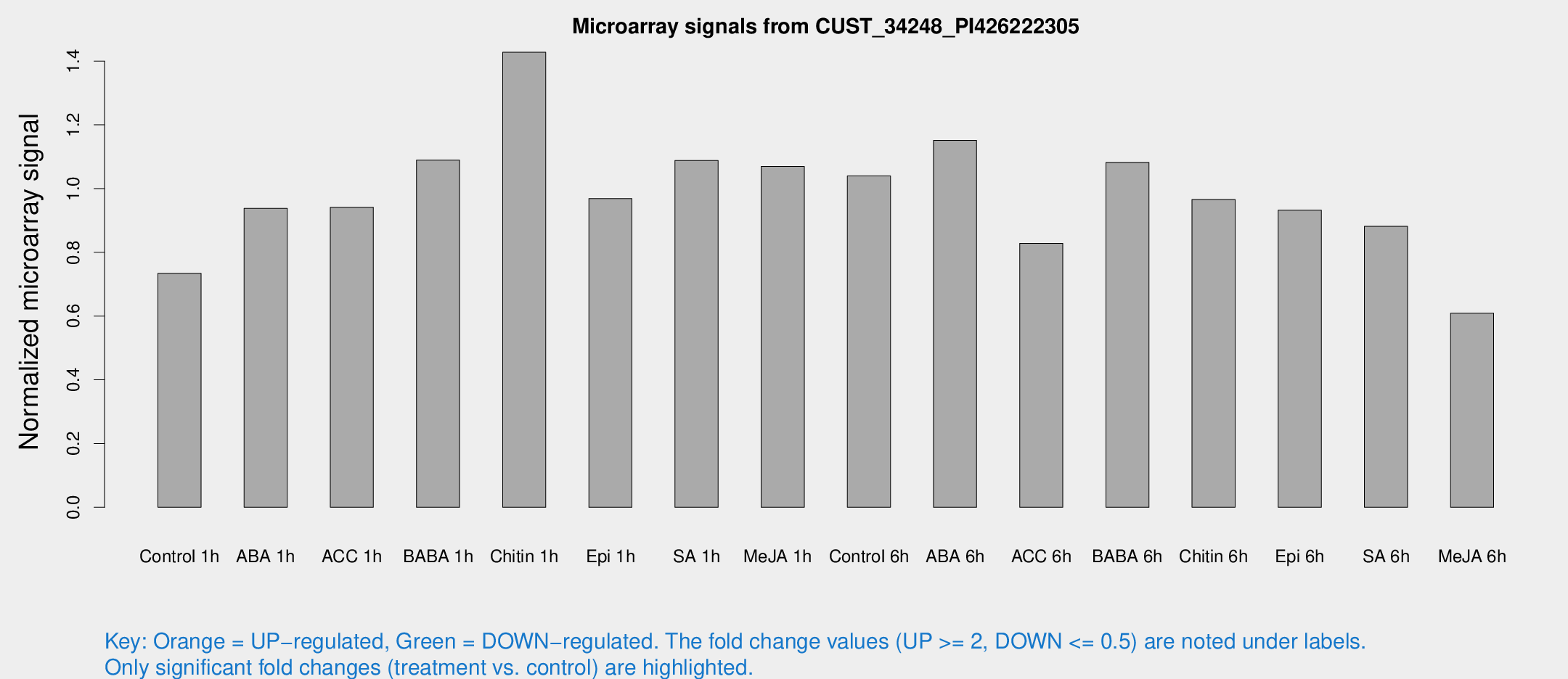
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4064.53 | 347.751 | 0.734094 | 0.0423866 |
| ABA 1h | 4651 | 668.314 | 0.938033 | 0.0705111 |
| ACC 1h | 5343.11 | 386.278 | 0.941541 | 0.0773204 |
| BABA 1h | 5977.95 | 1059.43 | 1.08934 | 0.102152 |
| Chitin 1h | 7197.46 | 1031.67 | 1.42789 | 0.208708 |
| Epi 1h | 4626.33 | 273.233 | 0.968672 | 0.0559299 |
| SA 1h | 6277.26 | 935.693 | 1.08821 | 0.093394 |
| Me-JA 1h | 4843.81 | 453.932 | 1.06919 | 0.0765158 |
| Control 6h | 6141.19 | 1432.38 | 1.03956 | 0.19696 |
| ABA 6h | 6813.76 | 778.951 | 1.15122 | 0.0664682 |
| ACC 6h | 5393.26 | 941.669 | 0.828405 | 0.0669965 |
| BABA 6h | 6668.65 | 385.505 | 1.08218 | 0.0624821 |
| Chitin 6h | 5706.17 | 544.906 | 0.965573 | 0.0686129 |
| Epi 6h | 5850.99 | 647.619 | 0.932485 | 0.0899547 |
| SA 6h | 5029.69 | 998.108 | 0.881824 | 0.0967434 |
| Me-JA 6h | 3628.39 | 954.197 | 0.609145 | 0.139072 |
Source Transcript PGSC0003DMT400044819 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G35790.1 | +3 | 0.0 | 683 | 331/379 (87%) | glucose-6-phosphate dehydrogenase 1 | chr5:13956879-13959686 REVERSE LENGTH=576 |
