Probe CUST_34222_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34222_PI426222305 | JHI_St_60k_v1 | DMT400030652 | GTCCTAATTCTGAGGAGGCCCCTACACATATGTAATTGTGTTTATCATTCTTGAAAATTA |
All Microarray Probes Designed to Gene DMG400011742
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34222_PI426222305 | JHI_St_60k_v1 | DMT400030652 | GTCCTAATTCTGAGGAGGCCCCTACACATATGTAATTGTGTTTATCATTCTTGAAAATTA |
Microarray Signals from CUST_34222_PI426222305
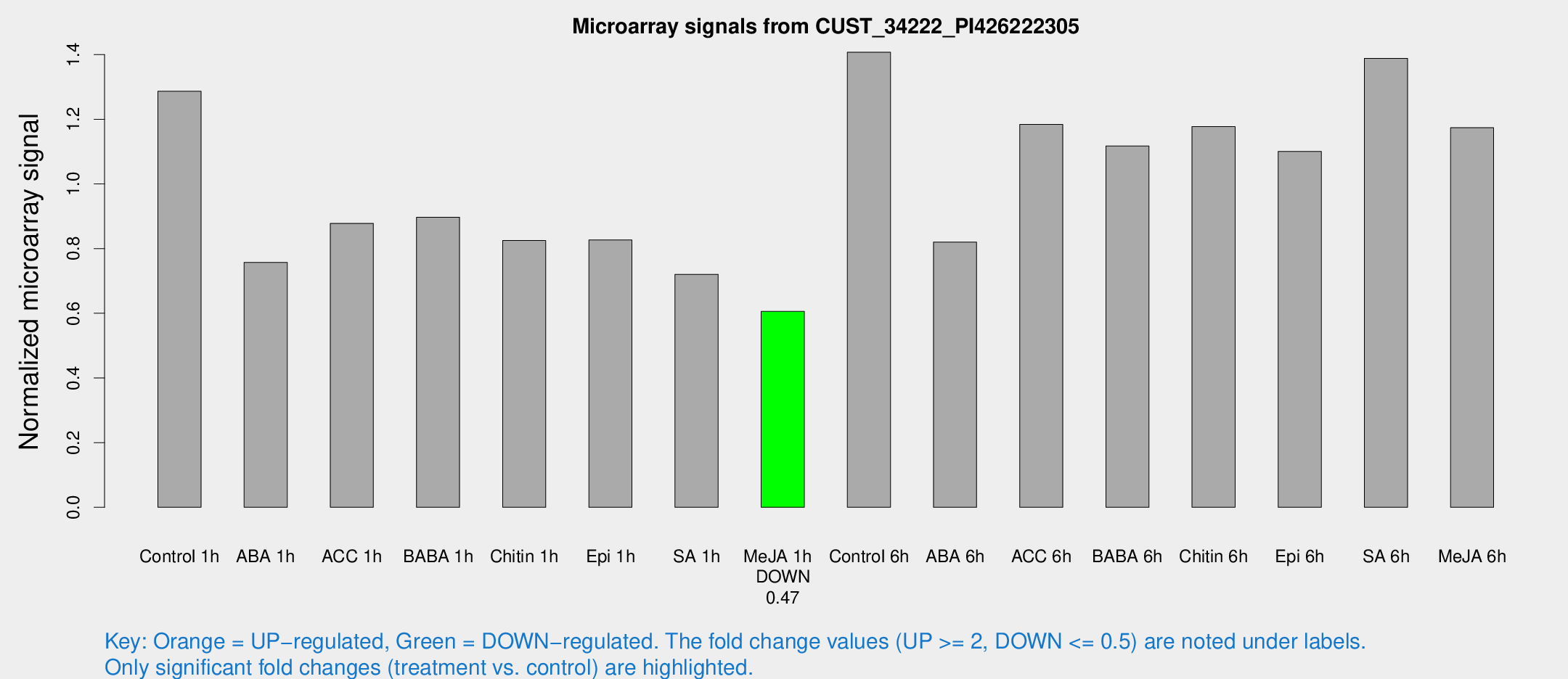
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1650.31 | 139.709 | 1.28659 | 0.0743303 |
| ABA 1h | 858.613 | 72.2363 | 0.75732 | 0.043828 |
| ACC 1h | 1192.64 | 215.277 | 0.878051 | 0.109117 |
| BABA 1h | 1137.92 | 193.741 | 0.896928 | 0.0758494 |
| Chitin 1h | 949.977 | 71.055 | 0.825026 | 0.0477326 |
| Epi 1h | 916.317 | 68.7317 | 0.826708 | 0.0552084 |
| SA 1h | 949.981 | 83.309 | 0.720534 | 0.0416862 |
| Me-JA 1h | 631.846 | 38.9358 | 0.605653 | 0.0351391 |
| Control 6h | 1854.08 | 332.205 | 1.40733 | 0.177079 |
| ABA 6h | 1118.14 | 95.9711 | 0.820383 | 0.0474501 |
| ACC 6h | 1744.99 | 161.221 | 1.18397 | 0.131579 |
| BABA 6h | 1601.06 | 122.206 | 1.11723 | 0.0757764 |
| Chitin 6h | 1597.9 | 92.483 | 1.17746 | 0.0680484 |
| Epi 6h | 1585 | 92.4344 | 1.1005 | 0.0712247 |
| SA 6h | 1770.27 | 194.787 | 1.3882 | 0.0802119 |
| Me-JA 6h | 1509.23 | 174.615 | 1.17382 | 0.0678388 |
Source Transcript PGSC0003DMT400030652 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G41920.1 | +2 | 8e-140 | 414 | 238/364 (65%) | GRAS family transcription factor | chr5:16779982-16781199 FORWARD LENGTH=405 |
