Probe CUST_34204_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34204_PI426222305 | JHI_St_60k_v1 | DMT400041149 | TGCTAAACACCCTTCAATTTTAAGAGAATCGATTAGTCCGTATCCATTCCAGTGTAGTCA |
All Microarray Probes Designed to Gene DMG400015925
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34186_PI426222305 | JHI_St_60k_v1 | DMT400041150 | TTTCTCTCGCAATTTATCTCATGGATTCTCAACACAAGAGTAATCGTCAACTCCAAAGAG |
| CUST_34204_PI426222305 | JHI_St_60k_v1 | DMT400041149 | TGCTAAACACCCTTCAATTTTAAGAGAATCGATTAGTCCGTATCCATTCCAGTGTAGTCA |
Microarray Signals from CUST_34204_PI426222305
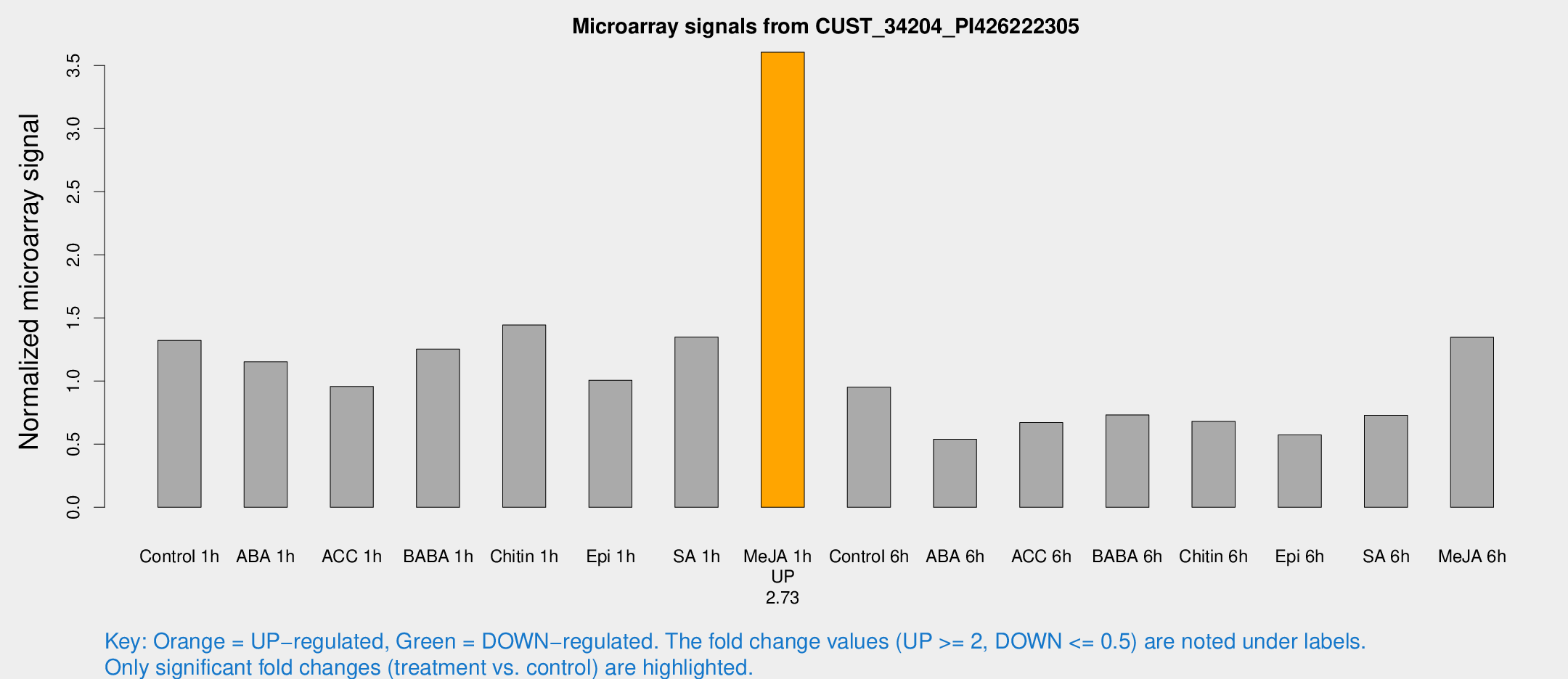
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 122.04 | 14.9556 | 1.32178 | 0.144455 |
| ABA 1h | 95.6645 | 17.2434 | 1.15207 | 0.124927 |
| ACC 1h | 97.8232 | 30.0965 | 0.956948 | 0.250792 |
| BABA 1h | 112.596 | 17.5567 | 1.25295 | 0.100129 |
| Chitin 1h | 119.103 | 11.4039 | 1.44375 | 0.204431 |
| Epi 1h | 79.2424 | 5.64027 | 1.00644 | 0.0715655 |
| SA 1h | 130.618 | 26.045 | 1.34802 | 0.281247 |
| Me-JA 1h | 270.252 | 27.3242 | 3.60515 | 0.213675 |
| Control 6h | 95.1771 | 26.2698 | 0.9514 | 0.222139 |
| ABA 6h | 54.392 | 11.3532 | 0.539201 | 0.0818236 |
| ACC 6h | 71.3615 | 9.76495 | 0.670855 | 0.0724844 |
| BABA 6h | 75.0735 | 6.26571 | 0.732108 | 0.0576815 |
| Chitin 6h | 66.1732 | 5.65654 | 0.681584 | 0.0582603 |
| Epi 6h | 67.0524 | 21.8 | 0.574112 | 0.284218 |
| SA 6h | 68.5473 | 13.9425 | 0.728928 | 0.0823654 |
| Me-JA 6h | 123.76 | 15.2899 | 1.34728 | 0.0899342 |
Source Transcript PGSC0003DMT400041149 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G43020.1 | +2 | 2e-52 | 182 | 83/102 (81%) | polyamine oxidase 2 | chr2:17891945-17894440 FORWARD LENGTH=490 |
