Probe CUST_33818_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33818_PI426222305 | JHI_St_60k_v1 | DMT400017004 | GATATTTCCTCAATTACGACTCTGCATTGCCCATCTGTTTGGGAGAGTAGCTTACCAAAT |
All Microarray Probes Designed to Gene DMG400006643
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33818_PI426222305 | JHI_St_60k_v1 | DMT400017004 | GATATTTCCTCAATTACGACTCTGCATTGCCCATCTGTTTGGGAGAGTAGCTTACCAAAT |
Microarray Signals from CUST_33818_PI426222305
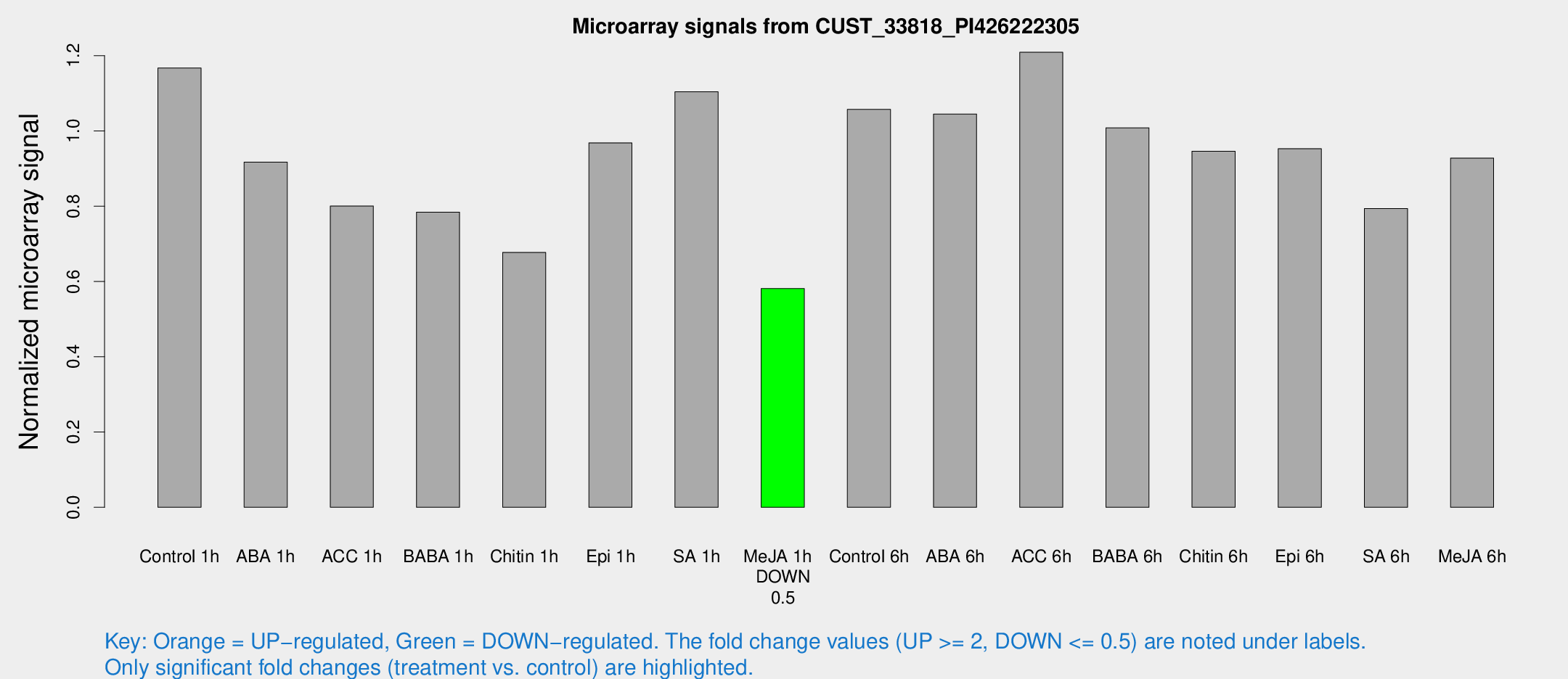
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 21055.3 | 1849.77 | 1.16733 | 0.0673958 |
| ABA 1h | 14564.1 | 843.283 | 0.916918 | 0.060175 |
| ACC 1h | 15921.3 | 3916.78 | 0.800423 | 0.180733 |
| BABA 1h | 14164.6 | 2815.05 | 0.784301 | 0.0941899 |
| Chitin 1h | 10965.1 | 806.27 | 0.677236 | 0.0391008 |
| Epi 1h | 15100.6 | 1215.35 | 0.968267 | 0.0559033 |
| SA 1h | 20509.3 | 2109.79 | 1.10397 | 0.135428 |
| Me-JA 1h | 8501.87 | 491.34 | 0.581103 | 0.065808 |
| Control 6h | 19434.8 | 2836.13 | 1.05713 | 0.0711925 |
| ABA 6h | 19964.1 | 1250.65 | 1.04486 | 0.0603254 |
| ACC 6h | 24964.6 | 1798.65 | 1.20914 | 0.113966 |
| BABA 6h | 20302.6 | 1370.62 | 1.00837 | 0.0698085 |
| Chitin 6h | 18069.2 | 1045.46 | 0.946035 | 0.0546197 |
| Epi 6h | 19300.4 | 1117.95 | 0.952935 | 0.107681 |
| SA 6h | 16744.4 | 5614.37 | 0.793717 | 0.296876 |
| Me-JA 6h | 16645.9 | 1328.3 | 0.927922 | 0.113275 |
Source Transcript PGSC0003DMT400017004 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G10985.1 | +1 | 8e-33 | 117 | 57/110 (52%) | senescence associated gene 20 | chr3:3442776-3443108 FORWARD LENGTH=110 |
