Probe CUST_33490_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33490_PI426222305 | JHI_St_60k_v1 | DMT400058271 | TCTCTCAACATTGTTCAGTATATTGATGAAAAATGGACTAATTCTGGTCCCTCTATTCTC |
All Microarray Probes Designed to Gene DMG400022624
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33490_PI426222305 | JHI_St_60k_v1 | DMT400058271 | TCTCTCAACATTGTTCAGTATATTGATGAAAAATGGACTAATTCTGGTCCCTCTATTCTC |
| CUST_33563_PI426222305 | JHI_St_60k_v1 | DMT400058270 | GAAGCTAGCCATGGAAACAAATTGTGTAAAGCTTCTAGGTTCATGGGCAAGCCCGTTTGT |
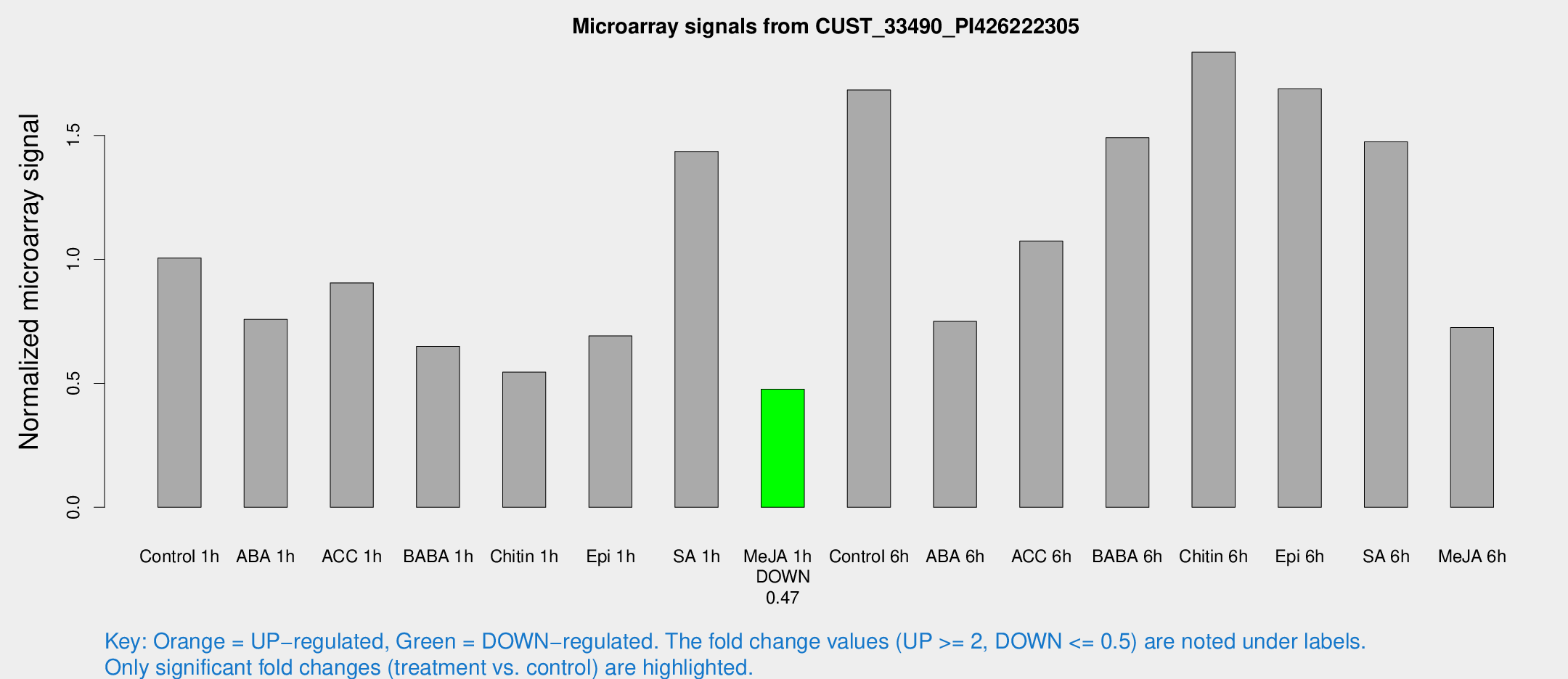
Microarray Signals from CUST_33490_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 199.215 | 31.3693 | 1.00581 | 0.102528 |
| ABA 1h | 130.58 | 12.8702 | 0.757944 | 0.0475044 |
| ACC 1h | 182.521 | 24.6676 | 0.905552 | 0.0752496 |
| BABA 1h | 125.755 | 23.4821 | 0.649229 | 0.0688162 |
| Chitin 1h | 94.6677 | 6.36818 | 0.545133 | 0.0444958 |
| Epi 1h | 115.738 | 7.44582 | 0.691449 | 0.0476734 |
| SA 1h | 298.373 | 68.491 | 1.4354 | 0.242482 |
| Me-JA 1h | 75.6579 | 6.87224 | 0.476717 | 0.0348443 |
| Control 6h | 336.584 | 57.4349 | 1.68347 | 0.170077 |
| ABA 6h | 153.83 | 9.5699 | 0.750023 | 0.0467485 |
| ACC 6h | 244.596 | 42.8327 | 1.07395 | 0.0645981 |
| BABA 6h | 327.28 | 42.8789 | 1.49113 | 0.206173 |
| Chitin 6h | 383.853 | 53.3836 | 1.83583 | 0.190186 |
| Epi 6h | 367.768 | 21.6297 | 1.68778 | 0.205775 |
| SA 6h | 307.416 | 79.3652 | 1.47413 | 0.301098 |
| Me-JA 6h | 164.202 | 55.2259 | 0.724983 | 0.289184 |
Source Transcript PGSC0003DMT400058271 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G10370.1 | +1 | 4e-89 | 269 | 141/227 (62%) | Glutathione S-transferase family protein | chr1:3397274-3398273 REVERSE LENGTH=227 |
