Probe CUST_33351_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33351_PI426222305 | JHI_St_60k_v1 | DMT400017852 | ATCAAAGAGGAATTTCGTCTTATTCTCAAGCCCGGTGGCTCGGTTGATTGCTAGTTGCTA |
All Microarray Probes Designed to Gene DMG400006933
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33351_PI426222305 | JHI_St_60k_v1 | DMT400017852 | ATCAAAGAGGAATTTCGTCTTATTCTCAAGCCCGGTGGCTCGGTTGATTGCTAGTTGCTA |
| CUST_33335_PI426222305 | JHI_St_60k_v1 | DMT400017851 | CAGCTGAAAGACATTGCGGATGGGCTTAGTGATGAGGAAGCTGATGAAGAAGATTCTGAT |
Microarray Signals from CUST_33351_PI426222305
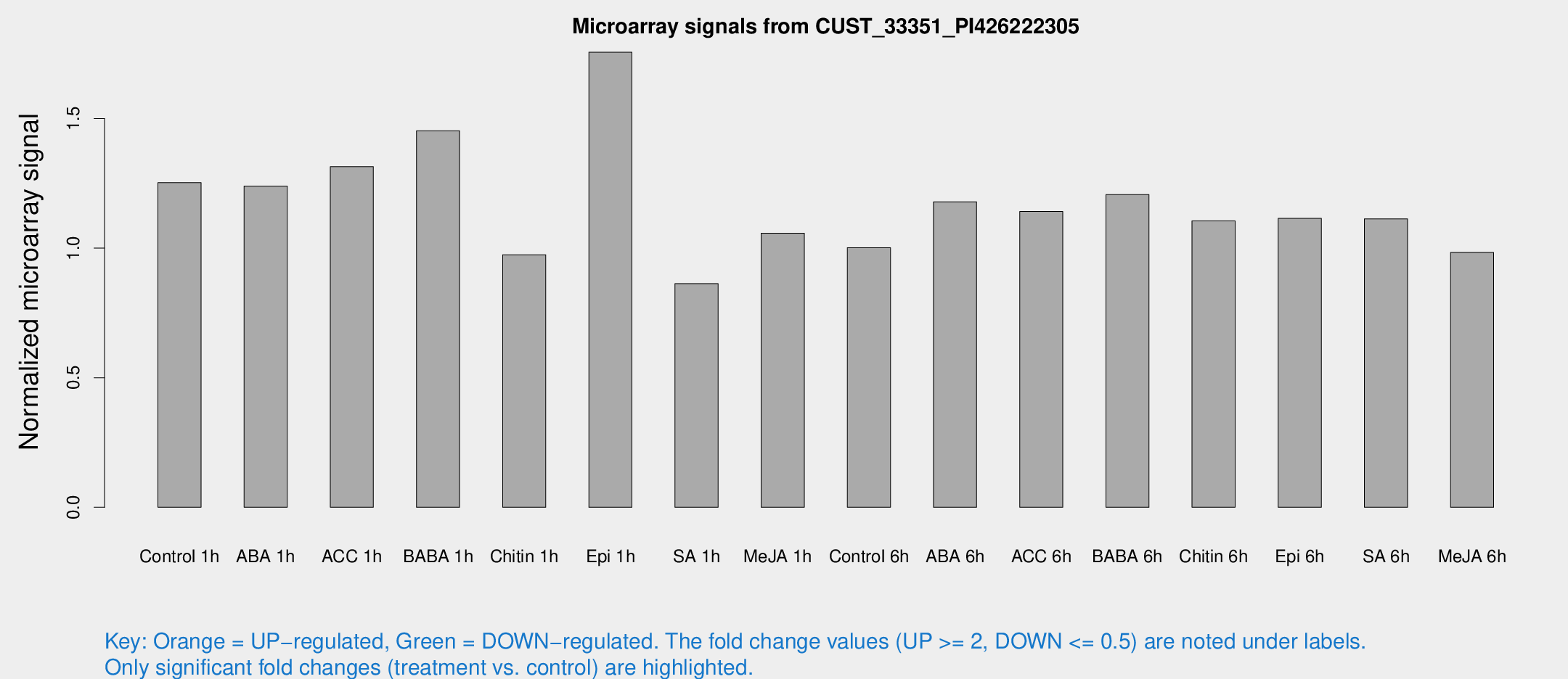
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.89197 | 4.52485 | 1.25265 | 0.618338 |
| ABA 1h | 7.67285 | 3.10528 | 1.2398 | 0.589724 |
| ACC 1h | 9.03976 | 3.50136 | 1.31412 | 0.547229 |
| BABA 1h | 11.2736 | 5.39556 | 1.45275 | 0.666581 |
| Chitin 1h | 5.72089 | 3.23519 | 0.974533 | 0.548164 |
| Epi 1h | 10.0247 | 3.26346 | 1.75593 | 0.58315 |
| SA 1h | 5.80231 | 3.20262 | 0.863187 | 0.478123 |
| Me-JA 1h | 5.64196 | 3.27298 | 1.0574 | 0.612247 |
| Control 6h | 6.69341 | 3.3247 | 1.00192 | 0.518274 |
| ABA 6h | 8.91322 | 3.46945 | 1.17876 | 0.536493 |
| ACC 6h | 8.88255 | 4.08986 | 1.1415 | 0.530805 |
| BABA 6h | 9.86148 | 3.73864 | 1.20679 | 0.569155 |
| Chitin 6h | 8.08767 | 3.6405 | 1.10514 | 0.547811 |
| Epi 6h | 8.66853 | 3.83933 | 1.11471 | 0.545097 |
| SA 6h | 7.26776 | 3.47503 | 1.11313 | 0.543137 |
| Me-JA 6h | 6.44407 | 3.29828 | 0.983228 | 0.514645 |
Source Transcript PGSC0003DMT400017852 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G18800.1 | +3 | 2e-104 | 315 | 158/222 (71%) | NAP1-related protein 2 | chr1:6481466-6483463 REVERSE LENGTH=256 |
