Probe CUST_3221_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3221_PI426222305 | JHI_St_60k_v1 | DMT400000653 | GTAGTAGCAAATTTTCCGTGACTGTCTATAGTAGTACACATTTTCTATCATAATCAAGGC |
All Microarray Probes Designed to Gene DMG400000232
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3221_PI426222305 | JHI_St_60k_v1 | DMT400000653 | GTAGTAGCAAATTTTCCGTGACTGTCTATAGTAGTACACATTTTCTATCATAATCAAGGC |
| CUST_3027_PI426222305 | JHI_St_60k_v1 | DMT400000652 | GTAGTAGCAAATTTTCCGTGACTGTCTATAGTAGTACACATTTTCTATCATAATCAAGGC |
Microarray Signals from CUST_3221_PI426222305
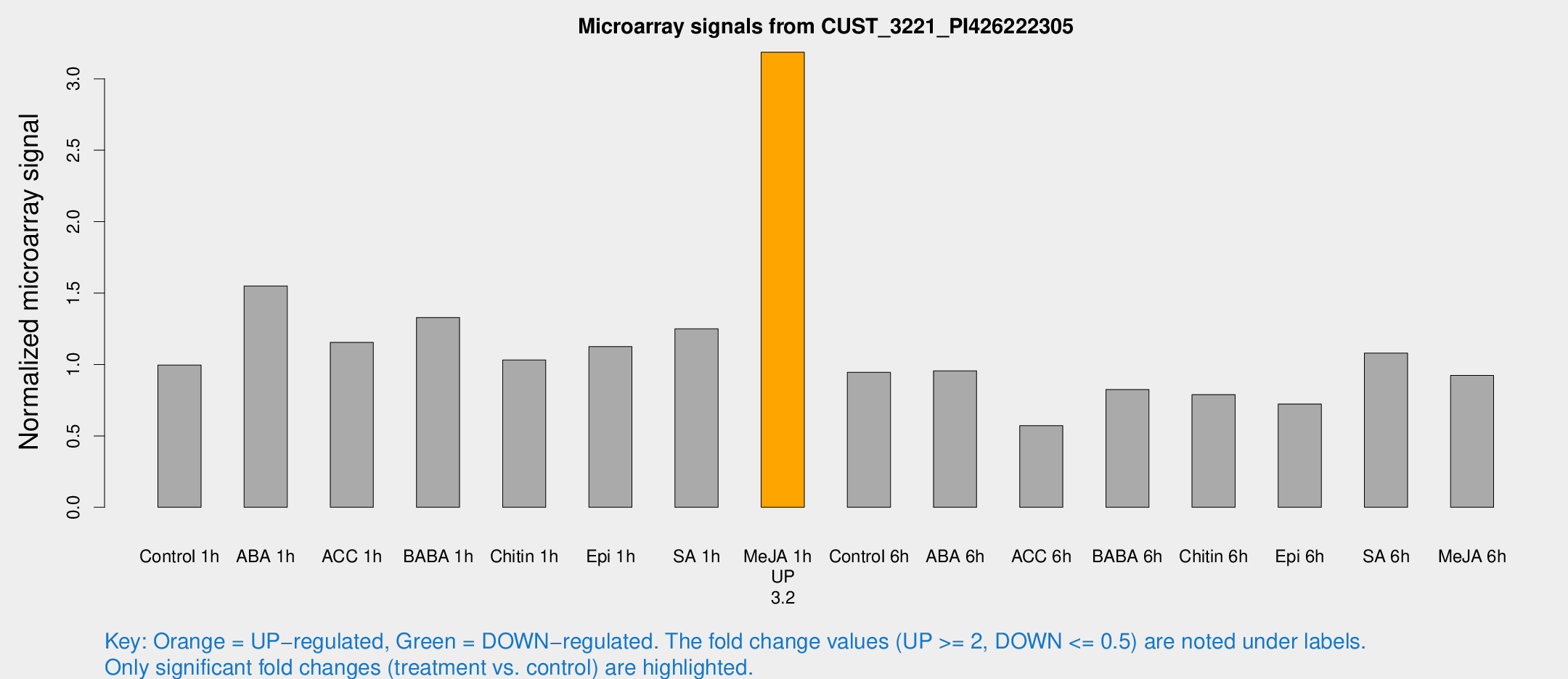
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 701.562 | 157.284 | 0.995403 | 0.232803 |
| ABA 1h | 962.407 | 221.854 | 1.55018 | 0.244725 |
| ACC 1h | 846.148 | 223.436 | 1.15404 | 0.249803 |
| BABA 1h | 871.278 | 121.468 | 1.32902 | 0.0865724 |
| Chitin 1h | 622.878 | 55.2748 | 1.03099 | 0.0792008 |
| Epi 1h | 673.795 | 119.682 | 1.12498 | 0.214443 |
| SA 1h | 857.413 | 49.6512 | 1.25049 | 0.0723304 |
| Me-JA 1h | 1759.66 | 219.328 | 3.18629 | 0.184049 |
| Control 6h | 650.193 | 103.144 | 0.945281 | 0.0796865 |
| ABA 6h | 705.369 | 145.031 | 0.955316 | 0.140678 |
| ACC 6h | 447.486 | 65.1196 | 0.571636 | 0.0923952 |
| BABA 6h | 618.183 | 45.4557 | 0.823931 | 0.0478073 |
| Chitin 6h | 560.515 | 32.5996 | 0.788538 | 0.0469599 |
| Epi 6h | 544.444 | 31.6918 | 0.723093 | 0.0501551 |
| SA 6h | 718.039 | 66.9658 | 1.07966 | 0.0824985 |
| Me-JA 6h | 614.314 | 35.6941 | 0.923163 | 0.053514 |
Source Transcript PGSC0003DMT400000653 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G01260.1 | +2 | 5e-73 | 252 | 120/173 (69%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr1:109595-111367 FORWARD LENGTH=590 |
