Probe CUST_31831_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31831_PI426222305 | JHI_St_60k_v1 | DMT400053121 | GGACAACTAAATAGGGACGTAGGGAGTAACATATAGTAGATTTTTTCTCAACTAATCTCT |
All Microarray Probes Designed to Gene DMG400020608
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31831_PI426222305 | JHI_St_60k_v1 | DMT400053121 | GGACAACTAAATAGGGACGTAGGGAGTAACATATAGTAGATTTTTTCTCAACTAATCTCT |
| CUST_31825_PI426222305 | JHI_St_60k_v1 | DMT400053122 | TCATGGATGAGAGAATCTGAAACAATGAATGATGGTTGTGCATGGAGAAAATATGGGCAG |
Microarray Signals from CUST_31831_PI426222305
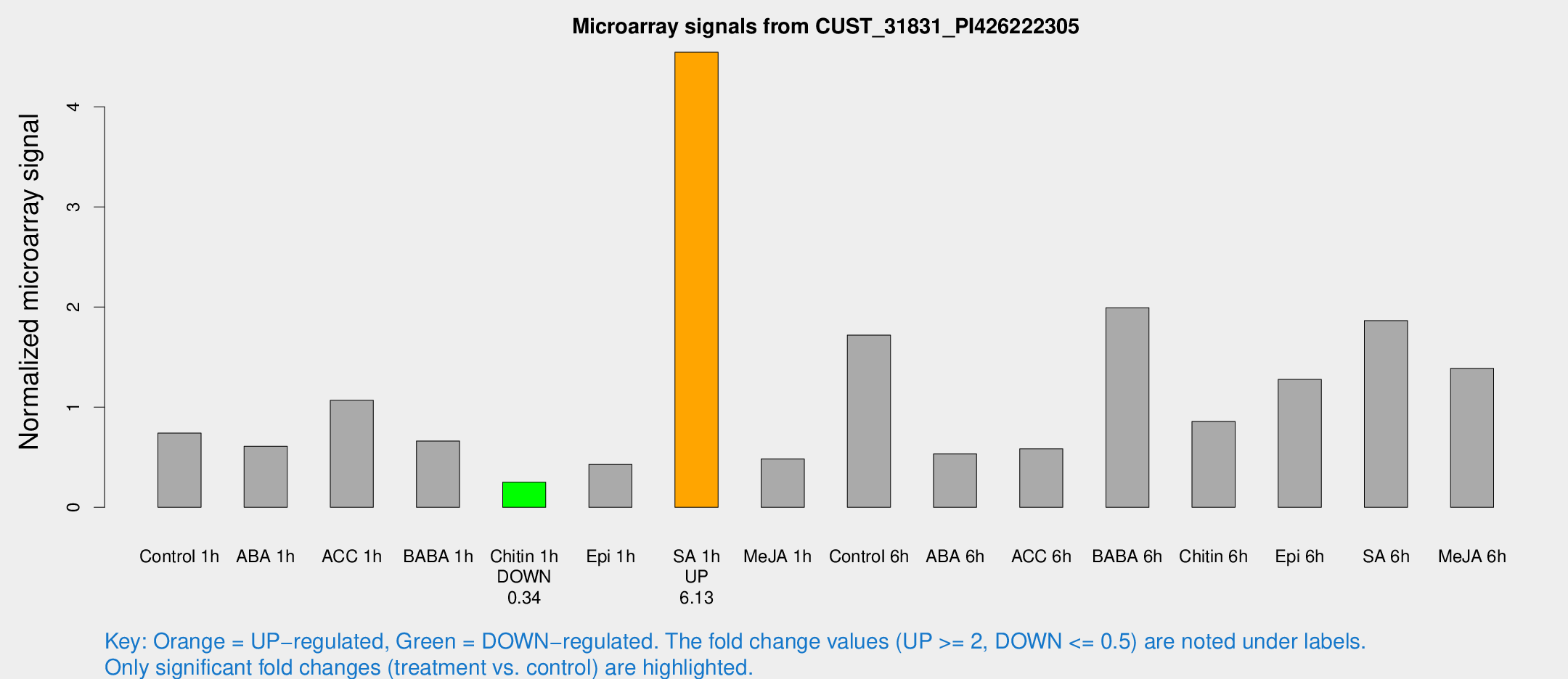
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 24.7365 | 7.80203 | 0.740998 | 0.179718 |
| ABA 1h | 21.1497 | 9.70132 | 0.609402 | 0.489485 |
| ACC 1h | 34.8542 | 6.95703 | 1.06897 | 0.264649 |
| BABA 1h | 22.2625 | 6.85665 | 0.66236 | 0.370486 |
| Chitin 1h | 6.9061 | 3.6384 | 0.251457 | 0.133575 |
| Epi 1h | 12.6043 | 3.63778 | 0.428815 | 0.163115 |
| SA 1h | 142.963 | 11.0242 | 4.54551 | 0.287121 |
| Me-JA 1h | 13.9695 | 5.34841 | 0.482216 | 0.25316 |
| Control 6h | 58.3074 | 17.0314 | 1.72044 | 0.464013 |
| ABA 6h | 21.112 | 7.52105 | 0.532325 | 0.289264 |
| ACC 6h | 22.7414 | 6.48427 | 0.584638 | 0.339806 |
| BABA 6h | 69.2192 | 9.05591 | 1.99377 | 0.214679 |
| Chitin 6h | 30.3018 | 8.48617 | 0.857765 | 0.263116 |
| Epi 6h | 46.978 | 11.8297 | 1.27735 | 0.431052 |
| SA 6h | 57.4752 | 8.30397 | 1.86502 | 0.497173 |
| Me-JA 6h | 47.6963 | 17.7953 | 1.38765 | 0.404985 |
Source Transcript PGSC0003DMT400053121 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G66560.1 | +3 | 4e-13 | 67 | 28/54 (52%) | WRKY DNA-binding protein 64 | chr1:24833579-24834631 FORWARD LENGTH=249 |
